Membrane Protein Structure Overview present and future
This overview explores the structure of membrane proteins, highlighting their crucial roles in transport and transduction processes across cellular membranes. Despite constituting about 20% of all proteins, membrane proteins remain less characterized than their water-soluble counterparts. The article discusses experimental challenges, current structural analyses, and advancements in crystallization and purification techniques. Relevant resources and research findings are provided to further understand the complexities and functions of these essential biomolecules.

Membrane Protein Structure Overview present and future
E N D
Presentation Transcript
Membrane Protein Structure Overview • present and future D.C. Rees Caltech/HHMI
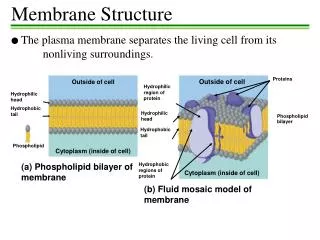
Membrane Proteins - Interests and Challenges participate in transport and transduction processes that mediate the flow of matter, energy and information across the membrane bilayer are poorly characterized relative to water-soluble proteins due to experimental challenges of mimicking the membrane and water-bilayer interfaces constitute an estimated ~20% of all proteins, yet only ~40 distinct structures are available membrane protein structure websites: http://blanco.biomol.uci.edu/Membrane_Proteins_xtal.html http://www.mpibp-frankfurt.mpg.de/michel/public/memprotstruct.html
membrane protein abundances 15-30% total, 10-20% > 2 TM Liu & Rost Prot. Sci.10, 1970 (2001)
Membrane protein structure: Status bioenergetic systems photosynthetic reaction centers (bacterial, PSI, PSII) bacteriorhodopsin (BR, HR, SR) respiratory complexes (II, III, IV and V) FDH, NarGHI light harvesting complexes ADP/ATP carrier transporters Ca+2 ATPase AcrB ABC transporters (MsbA, BtuCD) MFS (LacY, GlpT) channels/receptors K+ (KcsA, MthK, KirBac1.1, KvAP) mechanosensitive (MscL, MscS) aquaporins (GlpT, AQP) ClC AchR rhodopsin b-barrel 8,10,12,14,16,18,22 b-strands
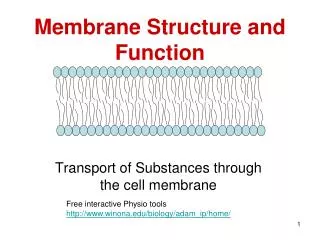
Structural analysis of membrane proteins natural sources or cloning & expression (recombinant) solubilization (detergents) purification crystallization structure determination
10-30 homologs membrane protein crystallization: approaches detergents/additives - RC lipidic phases - BR Fab complexes - cytochrome c oxidase proteolysis - KcsA sequence engineering - OmpX homologues - MscL
BtuCD structure determination 3.2 Å resolution Locher et al. Science 296, 1091 (2002)
BtuCD: molecular architecture of an ABC transporter two-fold NCS BtuC BtuC periplasm cytoplasm BtuC BtuD
surface polarity of membrane proteins reflects the solvent environment
surface hydrophobicity profile for membrane proteins ARBB31, 207 (2002) Nagle & Tristram-Nagle, COSB10, 474 (2000)
helix tilt and helix-helix crossing angle distributions average tilt = 23˚ + 10˚ ARBB31, 207 (2002)
Hydrophobicities of buried/surface residues membrane proteins vs water-soluble proteins TM region, 16 a-helical mps buried / surface Asp Ile +1 -1 0 hydrophobicity surface buried water-soluble proteins Eisenberg et al. hydrophobicity scale Faraday Symp. Chem. Soc.17, 109 (1982) Science 245, 510 (1989) ARBB31, 207 (2002)
Membrane protein - water-soluble protein comparison water membrane membrane proteins and water soluble proteins have similar interior apolarities packing densities surface areas stabilities (?) and differ in surface polarities helix-helix packing distributions tertiary folds what is the role of solvent in defining protein structures?
B.W. Matthews Ann. Rev. Phys. Chem.27, 493 (1976) http://www.mpibp-frankfurt.pg.de/ michel/public/memprotstruct.html photosynthetic reaction center myoglobin progress in membrane protein structure determinations parallels that of water-soluble proteins with a ~25 year offset
“Should we designate a certain percentage of targets to membrane proteins and/or protein complexes?” YES!!! Membrane protein challenges: overexpression of eukaryotic proteins solubilization (detergents) state-specific stabilizers
Acknowledgments (~t-20 years to present) RC: J. Allen, T. Yeates, A. Chirino, A. Yeh, H. Axelrod, G. Feher FRD: T. Iverson, C. Luna-Chavez, G. Cecchini MscL: G. Chang, R. Spencer, A. Lee, M. Barclay, R. Bass, P. Strop, J. Choe, S. Steinbacher MscS: R. Bass, P. Strop, M. Barclay, Y. Poon BtuCD: K. Locher, A. Lee, L. Borths general: R. Spencer, L. DeAntonio, D. Eisenberg