Chap. 8A Nucleotides and Nucleic Acids
350 likes | 579 Vues
Chap. 8A Nucleotides and Nucleic Acids. Some Basics Nucleic Acid Structure Nucleic Acid Chemistry Other Functions of Nucleotides. Fig. 8-12. X-ray diffraction pattern of DNA fibers. Intro. to Nucleotides and Nucleic Acids.

Chap. 8A Nucleotides and Nucleic Acids
E N D
Presentation Transcript
Chap. 8A Nucleotides and Nucleic Acids • Some Basics • Nucleic Acid Structure • Nucleic Acid Chemistry • Other Functions of Nucleotides Fig. 8-12. X-ray diffraction pattern of DNA fibers.
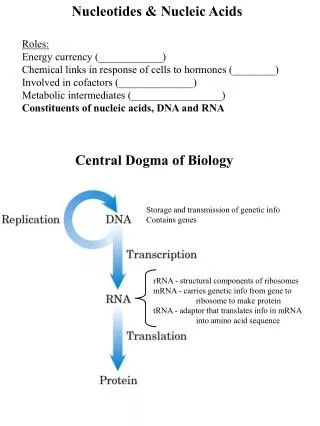
Intro. to Nucleotides and Nucleic Acids Nucleotides have a variety of roles in cellular metabolism. They are the energy currency in metabolic reactions, the essential chemical links in the response of cells to hormones and other stimuli, and the structural components of a variety of enzyme cofactors and metabolic intermediates. They are also constituents of the nucleic acids, deoxyribonucleic acid (DNA) and ribonucleic acid (RNA). The amino acid sequence of every protein in a cell, and the nucleotide sequence of every RNA, is specified by a nucleotide sequence in genomic DNA. Segments of DNA specifying the synthesis of a functional protein or RNA product are called genes. The storage and transmission of biological information are the only known functions of DNA. RNAs have a broader range of functions. Ribosomal RNAs (rRNAs) are structural and catalytic components of ribosomes. Messenger RNA (mRNAs) carry genetic information specifying the sequences of proteins. Transfer RNAs (tRNAs) are adaptor molecules that participate in translating the information in mRNA into a specific sequence of amino acids in a polypeptide. There is also a wide variety of special-function RNAs, including some called ribozymes that have enzymatic activity.
Structure of Nucleotides Nucleotides contain three components: 1) a nitrogen-containing base, 2) a pentose, and 3) one or more phosphates (Fig. 8-1a). In the absence of the phosphate group(s), the molecule is called a nucleoside. The nitrogenous bases are derivatives of pyrimidine and purine. The numbering of the ring atoms of pyrimidines and purines is illustrated in Fig. 8-1b. The numbering of the pentose rings follows the convention outlined in Chap. 7, except that the carbon numbers in the pentoses of nucleotides and nucleosides are given a prime (‘) designation to distinguish them from the numbered atoms of the bases. The base of a nucleotide is joined covalently (at N-1 of pyrimidines and N-9 of purines) in an N-ß-glycosyl bond to the 1’ carbon of the pentose, and the phosphate is esterified commonly to the 5’ carbon. Water is removed in the formation of the N-ß-glycosyl bond as occurs in O-glycosidic bond formation.
Major Bases of Nucleic Acids Both DNA and RNA contain two major purine bases, adenine (A) and guanine (G). Both nucleic acids also contain the pyrimidine, cytosine (C), and a second pyrimidine that is thymine (T) in DNA and uracil (U) in RNA. Only occasionally does thymine occur in RNA or uracil in DNA. In some cases the names of the bases reflect the sources from which they originally were isolated. Guanine, for example, was first isolated from guano (bird manure) and thymine was first isolated from thymus tissue.
Nomenclature of Nucleosides & Nucleotides The names of the nucleosides and nucleotides containing the five common bases are listed in Table 8-1.
The Pentoses of Nucleotides Nucleotides have two kinds of pentoses. The recurring deoxyribonucleotide units of DNA contain 2’-deoxy-D-ribose, and the ribonucleotide units of RNA contain D-ribose. In both types of nucleotides the pentoses exist in their ß-furanose (closed five-membered ring) forms. The formation of the ß-D-ribofuranose ring from the straight-chain aldehyde form of D-ribose in solution is illustrated in Fig. 8-3a. Deoxyribose undergoes a similar interconverion in solution, but in DNA exists solely as ß-2’-deoxy-D-ribofuranose. Deoxyribofuranose and ribofuranose rings in nucleotides exist in four different puckered conformations (Fig. 8-3b). In all cases, four of the five ring atoms are nearly in the same plane. The fifth atom (C-2’ or C-3’) is on either the same (endo) or the opposite (exo) side of the plane relative to the C-5’ atom.
Deoxyribonucleotides of DNA The structures and names of the four major deoxyribonucleotides (deoxyribonucleoside 5’-monophosphates) of DNA are shown below (Fig. 8-4a). All nucleotides are shown in their free form at pH 7.0. The deoxyribonucleotide units of DNA are usually symbolized as A, G, T, and C, and sometimes as dA, dG, dT, and dC. In their free forms, the deoxyribonucleotides are commonly abbreviated dAMP, dGMP, cTMP, and dCMP. For each nucleotide in the figure, the more common name is followed by the complete name in parentheses. All abbreviations assume that the phosphate group is at the 5’ position. The nucleoside portion of each molecule is shaded in light red.
Ribonucleotides of RNA The structures and names of the four major ribonucleotides (ribonucleoside 5’-monophosphates) of RNA are shown below (Fig. 8-4b). All nucleotides are shown in their free form at pH 7.0. The ribonucleotide units of RNA are usually symbolized as A, G, U, and C. In their free forms, the ribonucleotides are commonly abbreviated AMP, GMP, UMP, and CMP. For each nucleotide in the figure, the more common name is followed by the complete name in parentheses. All abbreviations assume that the phosphate group is at the 5’ position. The nucleoside portion of each molecule is shaded in light red.
Minor Bases Both DNA and RNA also contain some minor bases (Fig. 8-5). In DNA, the most common of these are methylated forms of the major bases (Panel a). Minor bases of many types occur in RNAs (Panel b). If the modification occurs directly on one of the ring atoms of the pyrimidine or purine base, the convention is to simply indicate the ring position of the substituent by its number, e.g., 5-methylcytidine. The convention changes when the substituent atom is exocyclic (not within the ring structure). In this case the type of atom is identified and the ring position to which it is attached is denoted with a superscript, e.g., N6-methyladenosine.
Some Adenosine Monophosphates Cells also contain nucleotides with phosphate groups in positions other than on the 5’ carbon of the pentose ring (Fig. 8-6). For example, ribonucleoside 2’,3’-cyclic monophosphates are isolatable intermediates, and ribonucleoside 3’-monophosphates are end products of the hydrolysis of RNA by certain ribonucleases. Other variants are adenosine 3’,5’-cyclic monophosphate (cAMP), which is a very important molecule in some signal transduction pathways.
Phosphodiester Linkages in the Covalent Backbone of DNA and RNA The successive nucleotides in DNA and RNA are covalently linked through phosphate-group bridges in which the 5’-phosphate of one nucleotide unit is joined to the 3’-hydroxyl group of the next, creating a phosphodiester linkage (Fig. 8-7). Thus, the covalent backbones of nucleic acids consist of alternating phosphate and pentose residues, and the nitrogenous bases may be regarded as side groups joined to the backbone at regular intervals. The backbones of both DNA and RNA are hydrophilic. The hydroxyl groups of the sugar residues form hydrogen bonds with water. The phosphate groups, with a pKa near 0, are completely ionized and negatively charged at pH 7. The negative charges are generally neutralized by ionic interactions with positive charges on proteins, metal ions, and polyamines. All the phosphodiester linkages in DNA and RNA have the same orientation along the chain giving each linear nucleic acid strand a specific polarity and distinct 5’ and 3’ ends. By definition, the 5’ end lacks a nucleotide at the 5’ position and the 3’ end lacks a nucleotide at the 3’ position.
Hydrolysis of RNA by Alkali The covalent backbone of DNA and RNA is subject to slow, nonenzymatic hydrolysis of its phosphodiester bonds. In vitro, RNA is hydrolyzed rapidly under alkaline conditions, but DNA is not. This is because the 2’-hydroxyl group in the ribose moieties of RNA is directly involved in the cleavage process (Fig. 8-8). 2’,3’-cyclic monophosphate nucleotides are the first products of the action of alkali on RNA and are subsequently hydrolyzed further to yield a mixture of 2’- and 3’-nucleoside monophosphates.
Representations of Nucleotides The nucleotide sequences of nucleic acids can be represented as illustrated below for a segment of DNA with five nucleotide units. The phosphate groups are symbolized by circled Ps, and each deoxyribose is symbolized by a vertical line, from C-1’ at the top to C-5’ at the bottom. The connecting lines between nucleotides (which pass through the P symbols) are drawn diagonally from the middle (C-3’) of the deoxyribose of one nucleotide to the bottom (C-5’) of the next. Some simpler representations of this pentadeoxyribonucleotide are pA-C-G-T-AOH, pApCpGpTpA, and pACGTA. Note that the sequence of a single strand of nucleic acid is always written with the 5’ end at the left and the 3’ end at the right, that is in the 5’3’ direction. A short nucleic acid such as shown in the figure is referred to as an oligonucleotide. This term is generally applied to nucleotides of 50 or fewer residues. Longer nucleic acids are referred to as polynucleotides.
Tautomeric Forms of Uracil Free pyrimidine and purine bases may exist in two or more tautomeric forms depending on the pH. Uracil, for example, occurs in lactam, lactim, and double lactim forms depending on the pH (Fig. 8-9). Certain tautomeric forms predominate at neutral pH, and these are the structures shown for the five common purines and pyrimidines in Fig. 8-2. These are the tautomers that are present in the bases in DNA and RNA.
Nucleotide Absorption Spectra All nucleotide bases absorb UV light, and nucleic acids are characterized by a strong absorption at wavelengths near 260 nm (Fig. 8-10). Plotted in this figure is the variation in molar extinction coefficient, , as a function of wavelength. The molar extinction coefficients at 260 nm are listed in the attached table. The spectra of corresponding ribonucleotides and deoxyribonucleotides are essentially identical. For mixtures of nucleotides, a wavelength of 260 nm is used for absorption measurements.
Watson and Crick Base-pairing in DNA The functional groups of pyrimidines and purines are ring nitrogens, carbonyl groups, and exocyclic amino groups. Hydrogen bonds involving the amino and carbonyl groups are the most important mode of interactions between two complementary strands of nucleic acid. The most common hydrogen-bonding patterns are those defined by Watson and Crick in 1953, in which A bonds specifically to T (or U) and G bonds to C (Fig. 8-11). These two types of base pairs predominate in double-stranded DNA and RNA, and the tautomers shown in Fig. 8-2 are responsible for these types of base pairs. It is this specific pairing of bases that permits the duplication of genetic information in DNA, as discussed below.
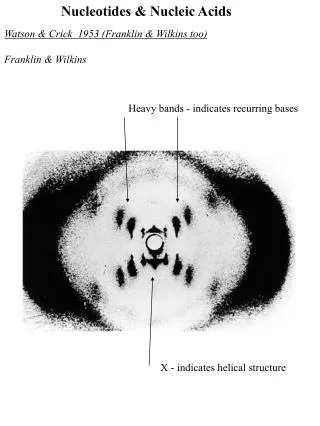
Watson-Crick Model for the Structure of Double-helical DNA (I) A model for the structure of DNA was proposed by Watson and Crick in 1953. Their model was based on a number of pieces of information that were available at the time about the composition of DNA and the x-ray diffraction properties of DNA fibers. Most importantly, x-ray diffraction studies of DNA fibers performed by Rosalind Franklin and Maurice Wilkins (Fig. 8-12) showed that DNA molecules are helical and exhibit two periodicities repeating along the length of the fiber--a primary repeat of 3.4 Å and a secondary repeat of 34 Å. In addition, Erwin Chargaff and colleagues had shown through DNA compositional analysis that the number of T residues always equals the number of A residues (A = T), and the number of G residues always equals the number of C residues (G = C). As a result, the sum of purine residues equals the sum of pyrimidine residues (A + G = T + C). Watson and Crick then set about to develop a structure that was consistent with these two sets of data (next slide).
Watson-Crick Model for the Structure of Double-helical DNA (II) In DNA, Watson and Crick proposed that two helical DNA chains are wound around the same axis to form a right-handed double helix (Fig. 8-13). They speculated that the two chains have an antiparallel orientation, and this was later proven to be true. The hydrophilic backbones of alternating deoxyribose and phosphate groups are on the outside of the helix facing the surrounding water. The furanose ring of each deoxyribose is in the C-2’ endo conformation. The purine and pyrimidine bases of both strands are stacked inside the double helix with their hydrophobic and nearly planar ring structures very close together and perpendicular to the axis of the helix. (Continued on the next slide).
Watson-Crick Model for the Structure of Double-helical DNA (III) Base stacking accounts for the 3.4 Å repeat along the length of the helix. The secondary repeat of about 34 Å was accounted for by the presence of 10 base pairs in each complete turn of the double helix. This was later modified to 10.5 base pairs per turn for DNA in aqueous solution. In the Watson-Crick model, A/T and G/C base pairing was proposed based on the fact that these combinations of bases fit well inside the double helix. Finally, the offset pairing of the two strands creates a major and a minor groove on the surface of the duplex. It should be noted that the double helix not only is stabilized by Watson-Crick base pairing between residues in the helix, but is also stabilized by base-stacking interactions that remove the bases from contact with water. The features of the double-helical model of DNA structure are supported by much chemical and biochemical evidence.
Double-helical Strand Complementarity The two antiparallel chains of double-helical DNA are not identical in either base sequence or composition. Instead, they are complementary to one another. Wherever adenine occurs in one chain, thymine occurs in the other. Similarly, guanine occurs opposite cytosine in the two chains (Fig.8-14).
Watson-Crick Model for DNA Replication The model for DNA structure immediately suggested to Watson and Crick a mechanism for the transmission of genetic information. The essential feature of the model is the complementarity of the two DNA strands in the double helix. As Watson and Crick were able to see, well before confirmatory data became available, this structure could logically be replicated by separating the two strands, and synthesizing a complementary strand for each (Fig. 8-15). Because nucleotides in each strand are joined in a sequence specified by the base-pairing rules stated above, each preexisting strand functions as a template to guide the synthesis of one complementary strand.
Structural Variation in DNA Double-helical DNA is a very flexible molecule. Considerable rotation is possible around several types of bonds in the phosphodeoxyribose backbone, and thermal fluctuation can produce bending, stretching, and unpairing (melting) of the strands. Many significant deviations from the Watson-Crick DNA structure are found in cellular DNA, some or all of which may be important in DNA metabolism. Note that these structural variations generally do not affect the fundamental properties of DNA, such as strand complementarity, antiparallel strands, and the requirement for A/T and G/C base pairs. Structural variations in DNA are due to three things: the different possible conformations of the deoxyribose, rotation about the contiguous bonds that make up the phosphodeoxyribose backbone (Fig. 8-16a), and free rotation about the C-1’-N-glycosyl bond (Fig. 8-16b). Because of steric constraints, purines in purine nucleotides are restricted to two stable conformations with respect to deoxyribose, called syn and anti (Fig. 8-16b). Pyrimidines generally are restricted to the anti conformation because of steric interference between the sugar and the carbonyl oxygen at C-2 of the pyrimidine.
The A, B, and Z Forms of DNA (I) The Watson-Crick structure of DNA is also referred to as B-form DNA, or B-DNA. B-DNA is the most stable structure for a random-sequence DNA molecule under physiological conditions and is therefore the standard structural reference in any study of the properties of DNA. Two structural variants that have been well characterized in crystal structures are the A- and Z-forms of DNA (Fig. 8-17). The properties of these forms are summarized in the Fig. 8-17 table (next slide). In general, A-DNA is a dehydrated form of DNA that may not occur in cells. A similar type of structure does occur in double helical RNA. There is evidence for some short tracts of Z-DNA in bacterial and eukaryotic cells. These Z-DNA tracts may play a role (as yet unidentified) in regulating the expression of some genes or in genetic recombination.
The A, B, and Z Forms of DNA (II) The DNA backbone of Z-DNA takes on a zigzag conformation. Certain nucleotide sequences fold into left-handed Z helices much more readily than others. Prominent examples are sequences in which pyrimidines alternate with purines, especially alternating C and G or 5-methyl-C and G residues. To form the left-handed helix in Z-DNA, the purine residues flip to the syn conformation, alternating with pyrimidines in the anti conformation.
DNA Palindromes and Mirror Repeats (I) It is thought that other sequence-dependent structural variations found in larger chromosomes may affect the function and metabolism of the DNA segments in their immediate vicinity. A rather common type of such DNA sequence is a palindrome. A palindrome is a word, phrase, or sentence that is spelled identically read either forward or backward. A example is the word ROTATOR. The term is applied to regions of DNA with inverted repeats of base sequence having twofold symmetry over two strands of DNA (Fig. 8-18). As shown in the next slide, such sequences have the potential to form unique DNA structures. When the inverted repeat occurs within each individual strand of the DNA, the sequence is called a mirror repeat. Mirror repeats cannot form the same types of structures as DNA palindromes.
DNA Palindromes and Mirror Repeats (II) DNA palindromes are self-complementary within each strand and therefore have the potential to form hairpin and cruciform (cross-shaped ) structures (Fig. 8-19). Sequences of these types are found in virtually every large DNA molecule and can encompass a few base pairs or thousands. The extent to which palindromes occur in cruciforms in cells is not known, although some cruciform structures have been demonstrated in vivo in E. coli. Self-complementary sequences cause isolated single strands of DNA (or RNA) in solution to fold into complex structures containing multiple hairpins.
Triple-helical DNA (I) Several unusual DNA structures involve three or four DNA strands. Nucleotides participating in a Watson-Crick base pair (Fig. 8-11) can form additional hydrogen bonds, particularly with functional groups in the major groove. For example, a cytidine residue if protonated can pair with a guanosine residue of a G/C base pair, and an thymidine can pair with the adenosine of an A/T pair (Fig. 8-20a). The N-7, O6, and N6 of purines are the atoms that participate in the hydrogen bonding of triplex DNA. These atoms are often referred to as Hoogsteen positions, and the non-Watson-Crick pairing is called Hoogsteen pairing after their discoverer, Karst Hoogsteen. This type of base pairing allows the formation of triplex DNAs.
Triple-helical DNA (II) Some triplex DNAs contain two pyrimidine strands and one purine strand. Others contain two purine strands and one pyrimidine strand. In Fig. 8-20b, triple-helical DNA containing two pyrimidine strands (red and white; sequence TTCCT) and one purine strand (blue; sequence AAGGAA) are shown. The blue and white strands are antiparallel and are paired by normal Watson-Crick base pairs. The third (all pyrimidine strand) is parallel to the purine strand and paired through non-Watson-Crick hydrogen bonds. The triplexes shown in Fig. 8-20a&b are most stable at low pH because the C/G.C+ triplet requires a protonated cytosine. In the triplex, the pKa of this cytosine is >7.5, which is raised considerably from its normal value of 4.2. Triplexes form most readily within long sequences containing only pyrimidines or only purines in a given strand.
Tetraplex DNA Four DNA strands can also pair to form a tetraplex (quadruplex), but this occurs readily only for DNA sequences with a very high proportion of guanosine residues (Fig. 8-20c&d). The guanosine tetraplex, or G tetraplex, is quite stable over a wide range of conditions. The orientation of strands in the tetraplex can vary as shown in Fig. 8-20e.
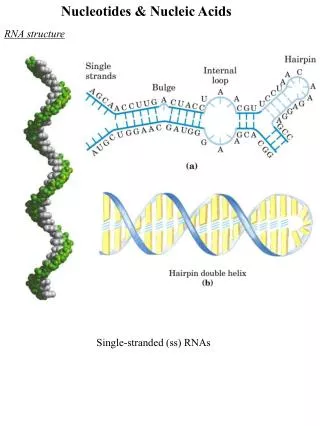
Intro. to RNA Structure As in the case of protein structure, it is sometimes useful to describe nucleic acid structure in terms of hierarchical levels of complexity (i.e., primary, secondary, and tertiary structure). The primary structure of a nucleic acid is its covalent structure and nucleotide sequence. Any regular, stable structure taken up by some or all of the nucleotides in a nucleic acid can be referred to as secondary structure. The complex folding of large chromosomes within eukaryotic chromatin and the bacterial nucleoid, or the elaborate folding of large tRNA or rRNA molecules, is generally considered tertiary structure. The process by which RNAs are formed on a DNA template is known as transcription. In bacteria, mRNAs can contain the protein coding sequences (red) for one gene (monocistronic) or multiple genes (polycistronic) (Fig. 8-21). mRNAs also contain noncoding regions (gray) at their ends and between protein coding sequences that may be involved in regulation of protein synthesis.
Base Stacking in Single-stranded RNA The product of transcription of DNA is always single-stranded RNA. The single strand tends to assume a right-handed helical conformation dominated by base-stacking interactions (Fig. 8-22). In this structure the bases are shown in yellow, the phosphorous atoms in orange, and the ribose and phosphate oxygens in green. Base stacking interactions are stronger between two purines than between a purine and pyrimidine or between two pyrimidines. The purine-purine interactions is so strong that a pyrimidine separating two purines is often displaced from the stacking pattern so that the purines can interact.
RNA Secondary Structures (I) Any self-complementary sequences in single-stranded RNAs produce more complex secondary structures (Fig. 8-23). Base pairing in RNA matches the pattern observed for DNA. Namely, G pairs with C, and A pairs with U (or the occasional T residue in some RNAs). One difference is that base pairing between G and U residues is allowed in RNA (Fig. 8-23a and Fig. 8-24, inset) when complementary sequences in two single strands of RNA pair with each other. The paired strands in RNA duplexes (or RNA-DNA duplexes) are antiparallel, as in double-helical DNA. The predominant double-stranded structure is an A-form right-handed double helix. The B-form double helix has not been observed for RNA. Strands of RNA that are precisely complementary over long regions of sequence are uncommon. Breaks in the regular A-form RNA double helix caused by mismatched or unmatched bases in one or both strands are common and result in bulges or internal loops (Fig. 8-23a). Hairpin loops form between nearly self-complementary (palindromic) sequences.
RNA Secondary Structures (II) The potential for base-paired helical segments in many RNAs is extensive, and the resulting hairpins are the most common type of secondary structure in RNA. The extensive secondary structure of an RNA is illustrated for the M1 RNA component of the enzyme RNase P of E. coli, that functions in the processing of tRNA (Fig. 8-24). This enzyme also contains a protein component (not shown). The two brackets in the figure indicate additional complementary sequences that may be paired in the three dimensional structure. The blue dots indicate non-Watson-Crick G/U base pairs (see inset). Note that G/U base pairs are only allowed when presynthesized RNA strand anneal together, and are not inserted by RNA polymerases into an RNA strand across from a G in a DNA template.
Three-dimensional Structure in RNA Extensive secondary structures, as illustrated for the M1 RNA, act as starting points for the folding of an RNA molecule into its precise three-dimensional structure. Other contributions are made by hydrogen bonds that are not part of standard Watson-Crick base pairs. These include bonds involving the 2’-hydroxyl group of ribose, and the oxygens of ribose phosphodiester bonds (Fig. 8-25a). The three-dimensional structures of phenylalanine tRNA of yeast, a hammerhead ribozyme from certain plant viruses, and the self-splicing intron of a mRNA from the ciliated protozoan, Tetrahymena thermophila are shown in the figure.