Transcription
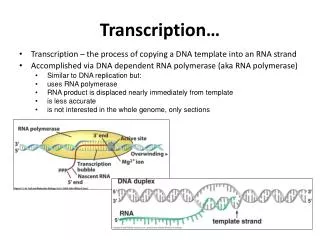
Transcription. Initiation. RNA polymerase binds to the promoter region, which is upstream of the gene that is being copied, and opens the double helix. Promoter region is high in A and T bases. Elongation. Begins building the single stranded mRNA in the 5’-3’ direction.

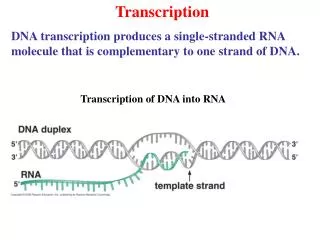
Transcription
E N D
Presentation Transcript
Initiation • RNA polymerase binds to the promoter region, which is upstream of the gene that is being copied, and opens the double helix. • Promoter region is high in A and T bases
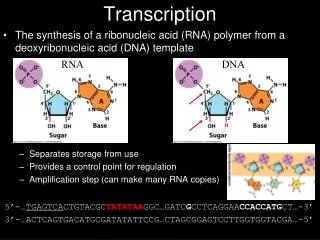
Elongation • Begins building the single stranded mRNA in the 5’-3’ direction. • No primer required. • One strand of the double stranded DNA acts as a template strand for RNA polymerase. • The strand not being used is the coding strand. • mRNA is a complement of the template strand, and an identical copy of the coding strand (with uracil).
Termination • Occurs when RNA polymerase reaches a terminator sequence at the end of the gene. • mRNA is released from the template.
Post-transcriptional Modifications • At this point, the mRNA (primary transcript) is not ready to leave the nucleus. A process known as capping and tailing must occur. • 5’ cap (7- methyl gunanosine) is added to the start of the ‘transcript’. • This protects the mRNA from digestion by nucleases and phosphatases as it exits the nucleus. • 3’ end gains approx. 200 adenine ribonucleotidescalled a poly-A tail by poly-A polymerase.
Further Modifications • DNA genes consist of coding regions: EXONS and non-coding regions: INTRONS • If the introns are translated, the protein will not fold properly and will be useless for the cell. • Spliceosomes remove intronsand join the remaining exons together so that coding regions are continuous. • Introns remain in the nucleus and are recycled. • Now it is mRNA transcript which is ready to be TRANSLATED by the ribosome.
Translation • mRNA has now exited the nucleus and is in the cytoplasm. • Initiation: the 5’ cap of the mRNA is recognized by ribosomes • Ribosomes: consist of 2 subunits; a large subunit (60s) and a small subunit (40s)
Elongation • The ribosome reads the mRNA 5’-3’ adding a new amino acid for every 3 nucleotides (codon). • Depending on where the ribosome starts reading, the reading frame will change. If the reading frame changes, the amino acids being coded for may also change.
tRNA • The molecule that delivers the amino acid is the transfer RNA. • tRNA is a small, single stranded nucleic acid resembling a clover. • The anti-codon (a series of three bases) recognizes the codon of the mRNA. • The opposite arm carries the corresponding amino acid. • Ex: mRNA codon = UAUanticodon arm= AUA
tRNA continued... • Only ONE amino acid is carried PER tRNA, therefore at least 20 tRNA’s are required (up to approx. 45). • With it’s amino acid tRNA is called aminoacyl-tRNA. • There are multiple codons for some amino acids: Serine: UCU, UCC, UCA, UCG. If the anticodon is: AGA, it can still bind to the codons UCC, UCA and/or UCG. • The third position (3’ on the mRNA and 5’ on the anticodon) can have non-standard base pairing: this is the ‘wobble hypothesis.’
Elongation • First codon recognized by the ribosome is the START codon (AUG)- methionine. Two sites: A (accepter site) P (peptide site) • The tRNA that carries methionine first enters the P site. • The next tRNA enters the A site. • Once the amino acids have been bonded (peptide bond), the tRNA exits the P site, and the tRNA from the A site takes its place. • The ribosome then shifts over one codon, and the new tRNA enters the A site.
Termination • The ribosome reaches a STOP codon • UGA, UAG, or UAA. • Protein called release factor aids in the release of the polypeptide chain from the ribosome. • The two subunits of the ribosome fall off of the mRNA. • Modifications to the polypeptide chain occur: Glycosylation- addition of a sugar group Phosphorylation- addition of a phosphate group