Plasmid Partitioning and Mobilization Mechanisms in Bacterial Cells
Explore the intricate mechanisms of parA and parB plasmid partitioning, as well as transcriptional repression and plasmid mobilization. Discover the roles of various proteins like DNA/RNA helicase and relaxase in plasmid segregation processes. Unravel the complex functions of topoisomerase/primase and type IV secretion systems in bacterial cell biology.

Plasmid Partitioning and Mobilization Mechanisms in Bacterial Cells
E N D
Presentation Transcript
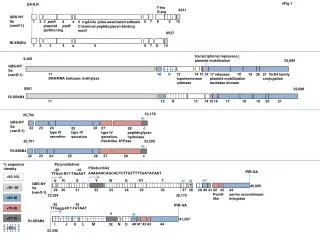
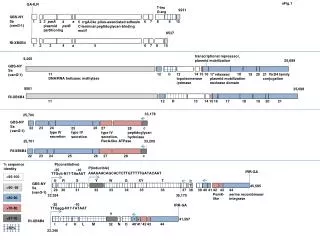
sFig.1 GA-ILR T-leu G-arg 9231 GBS-NY Sa (vanG-1) 1 2 a 6 7 8 9 10 3 parA plasmid partitioning 4 parB 5 rrgA-like pilus-associated adhesin C-terminal peptidoglycan-binding motif 8537 RI-XB6B4 1 2 3 4 a 5 6 7 8 10 transcriptional represssor, plasmid mobilization 9,265 25,689 GBS-NY Sa (vanG-1) 18 11 DNA/RNA helicase; methylase 12 b 14 15 16 21 VirD4 family conjugation 19 20 13 topoisomerase /primase 17 relaxase/ plasmid mobilization nuclease domain 8561 25,698 RI-XB6B4 11 12 H 13 14 15 16 17 18 19 20 21 33,178 25,704 GBS-NY Sa (vanG-1) 24 type IV secretion 22 23 26 25 type IV secretion 27 type IV secretion, RecA-like ATPase c 28 peptidoglycan hydrolase 33,209 25,701 RI-XB6B4 22 23 24 25 26 27 28 c % sequence identity P(constitutive) P(inducible) -35 -10 IRR-GA AAAAAACAGCACTCTTGTTTTTGATACAAT TTGctt-N17-TAaAAT >95-100 U R S Y W G XY T C T GBS-NY Sa (vanG-1) 45,585 >90- 95 40 29 30 31 32 33 34 35 36 37 38 39 41 43 42 PemK- like 44 serine recombinase/ integrase 33,354 39,170 >80-90 -35 -10 IRR-GA >70-80 TTGagg-N17-TATAAT Y >57-70 41,597 * * RI-XB6B4 I J K L M 32 N O 40’ 41’ 42 43 44 <30% 33,346