Advanced Labeling Kits for Accurate Gene Expression Profiling
Introducing three new labeling kits and protocols for accurate gene expression profiling. These kits provide enhanced signal intensity, reduce background noise, and detect messages missed by conventional methods. They offer advantages such as minimization of endogenous RNA priming and improved representation of expression profiles. Learn about the RT-Labeling Enzyme and the TrueLabeling-RT protocols that optimize experiments using GEArray kits. The AmpoLabeling-LPR kit amplifies signal intensity for low abundance messages. Explore how these protocols work and their benefits for gene-specific priming and expression profiling.

Advanced Labeling Kits for Accurate Gene Expression Profiling
E N D
Presentation Transcript
SuperArray Introduces Labeling Protocols Accessory Products for GEArray Original, Q, and S Series Kits
Three New Labeling Kits and Protocols • RT-Labeling Enzyme (RT) L-01 • Conventional reverse transcriptase and RNase inhibitor set • TrueLabeling-RT (TL-RT) L-02 • Reduces high background and “false positive” signals by minimizing endogenous priming of RNA • AmpoLabeling-LPR (LPR) L-03(N) • Amplifies the signal intensity of limiting amounts of total RNAor low abundance messages • Detects message ordinarily missed by conventional methods • Represents expression profile more accurately
Three New Labeling Kits and Protocols • RT-Labeling Enzyme (RT) L-01 • Conventional reverse transcriptase and RNase inhibitor set • TrueLabeling-RT (TL-RT) L-02 • Reduces high background and “false positive” signals by minimizing endogenous priming of RNA • AmpoLabeling-LPR (LPR) L-03(N) • Amplifies the signal intensity of limiting amounts of total RNAor low abundance messages • Detects message ordinarily missed by conventional methods • Represents expression profile more accurately
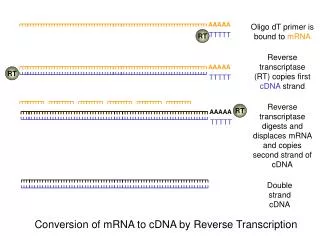
Gene-Specific Primers (A) 70 °C, 3 min 42 °C, 2 min Step 1: Annealing Mixture 5’ 3’ Total RNA RNase Inhibitor (RI) Reverse Transcriptase (RE) 42 °C, 25 or 90 min 95 °C, 5 min Step 2: Reverse Transcriptase Reaction 5’ 3’ Total RNA 3’ 5’ probe How Does the RT-Labeling Enzyme Protocol Work?
Advantages of RT-Labeling Enzyme • Experiments optimized previously using GEArray Original or Q Series Kits • Least expensive
Three New Labeling Kits and Protocols • RT-Labeling Enzyme (RT) L-01 • Conventional reverse transcriptase and RNase inhibitor set • TrueLabeling-RT (TL-RT) L-02 • Reduces high background and “false positive” signals by minimizing endogenous priming of RNA • AmpoLabeling-LPR (LPR) L-03(N) • Amplifies the signal intensity of limiting amounts of total RNAor low abundance messages • Detects message ordinarily missed by conventional methods • Represents expression profile more accurately
5’ 3’ Total RNA Gene-Specific primers (A) 70 °C, 3 min 42 °C, 2 min Step 1: Annealing Mixture 5’ 3’ Total RNA RNase Inhibitor (RI) TrueLabeling Reverse Transcriptase (AE) 42 °C, 90 min 95 °C, 5 min Step 2: Reverse Transcriptase Reaction 3’ 5’ probe How does the TrueLabeling-RT Enzyme Protocol Work?
Primers Enzyme - Conventional + - TrueLabeling + TrueLabeling Minimizes Endogenous RNA Priming 1:2 Serial Dilutions of cDNA probe
Gene-specific primers - + Conventional TrueLabeling Endogenous RNA Priming Causes High Background and a Skewed Expression Profile Human TGFb/BMP Signaling Pathway GEArray Q Series (HS-023) Human Universal Reference Total RNA
Advantages of TrueLabeling-RT • Minimizes endogenous RNA priming • Lowers background • Reduces possible false positive signals
Three New Labeling Kits and Protocols • RT-Labeling Enzyme (RT) L-01 • Conventional reverse transcriptase and RNase inhibitor set • TrueLabeling-RT (TL-RT) L-02 • Reduces high background and “false positive” signals by minimizing endogenous priming of RNA • AmpoLabeling-LPR (LPR) L-03(N) • Amplifies the signal intensity of limiting amounts of total RNAor low abundance messages • Detects message ordinarily missed by conventional methods • Represents expression profile more accurately
VS conventional labeling LPR 30 cycles AmpoLabeling-LPR Human TGFb/BMP Signaling Pathway GEArray Q Series (HS-023) 3.0 mg Human Universal Reference Total RNA
5’ 3’ Total RNA Random Primers (P) 70 °C, 3 min 37 °C, 10 min 5’ 3’ Total RNA How Does the AmpoLabeling-LPR Labeling Protocol Work? Step 1: Annealing Mixture +
5’ 3’ mRNA RNase Inhibitor (RI) Reverse Transcriptase (RE) 37 °C, 25 min 85 °C, 5 min 3’ 5’ cDNA RE How Does the AmpoLabeling-LPR Labeling Protocol Work? Step 2: RT Reaction
3’ 5’ cDNA Gene-specific primers (AF) Thermo-stable DNA Polymerase (LE) 85 °C, 5 min (85 °C, 1 min; 50 °C, 1 min., 72 °C, 1 min.) x 30 72 °C, 5 min 5’ 3’ probe How Does the AmpoLabeling-LPR Labeling Protocol Work? Step 3: Linear Polymerase Replication
30 cycles of LPR ID4 mg RNA 0.75 3.0 1.5 0.1 0.4 LPR Signal Is Proportional to Input Total RNA:As Little as 0.1 mg of Total RNA Detectable Human TGFb/BMP Signaling Pathway GEArray Q Series (HS-023) Human Universal Reference Total RNA
background corrected signal intensity mg RNA LPR Signal is Proportional to Input Total RNA
20 25 10 30 15 JUN cycles of LPR LPR Signal Is Proportional to Cycle Number:No More Than 30 Cycles are Required Human TGFb/BMP Signaling Pathway GEArray Q Series (HS-023) 3.0 mg Human Universal Reference Total RNA
LPR Signal Is Proportional to Cycle Number background corrected signal intensity cycles
RT-PCR conventional LPR ACVR1,2 COL3A1 INHA JUN, MADH2 TGFBR2 ACTB AmpoLabeling-LPR Represents Gene Expression Profiles More Accurately Human TGFb/BMP Signaling Pathway GEArray Q Series (HS-023) Human Universal Reference Total RNA
RT-PCR LPR G6PC AGP-1B PKCb UCP2 LIVER THYMUS LIVER THYMUS AmpoLabeling-LPR Represents Gene Expression Profiles More Accurately Mouse Insulin Signaling Pathway GEArray Q Series (MM-030) Mouse Liver or Thymus RNA
conventional LPR RT-PCR LIVER THYMUS AmpoLabeling-LPR Represents Gene Expression Profiles More Accurately Mouse Insulin Signaling Pathway GEArray Q Series (MM-030) Mouse Liver or Thymus RNA
Log (Liver:Thymus) for RT-PCR Log (Liver:Thymus) for conventional method Comparing Relative Gene Expression ProfilesObtained by RT-PCR and the Conventional Method
Log (Liver:Thymus) for RT-PCR Log (Liver:Thymus) for AmpoLabeling-LPR Comparing Relative Gene Expression ProfilesObtained by RT-PCR and AmpoLabeling-LPR
conventional 30 cycles LPR AKT1 FOS ACTB VS PDK1 LPR Detects Low Abundance Messages Ordinarily Missed by Conventional Methods GEArray Original Series Human PI-3 Kinase & AKT (hGEA9912030) 7.5 mg Human Universal Reference Total RNA Chemiluminescent Detection
conventional LPR Acaca Akt3 Bcl2l Cebpb G6pc VS Pdpk1 LPR Detects Low Abundance Messages Ordinarily Missed by Conventional Methods GEArray Q Series Mouse Insulin Signaling Pathway (MM-030) Mouse Universal Reference Total RNA Radioactive Detection
AmpoLabeling-LPR conventional VS LPR Detects Low Abundance Messages Ordinarily Missed by Conventional Methods Luo, Y., Cai, J., Ginis, I., Rao, M. (2003) Manuscript in Preparation GEArray S Series Mouse Stem Cell (MM-601.1) 0.6 mg Mouse D3 ES cell and 0.2 mg adult rat hippocampus total RNA Chemiluminescent Detection
Advantages of AmpoLabeling-LPR • Detects lower abundance messages • Detects message ordinarily missed by conventional methods • Represents gene expression profile more accurately • Use smaller amounts of input total RNA • Minimizes problems due to endogenous RNA priming • Lowers background • Reduces possible false positive (and false negative) signals
Three New Labeling Kits and Protocols • RT-Labeling Enzyme (RT) L-01 $ 50 • Includes: RI, RE, B, BN, C, D, E, H2O • TrueLabeling-RT (TL-RT) L-02 $ 100 • Includes: RI, AE, B, BN, C, D, E, H2O • AmpoLabeling-LPR (LPR) L-03(N) $ 200 • Includes: P, RI, RE, L, LE, BL, BN, C, H2O)
Acknowledgements • Ying Han • Kun Lu • Xiao Zeng THANK YOU.