cDNA Library
cDNA Library. MB206. Making a DNA library . Types of Libraries. (3’UTR). (5’UTR). Genomic DNA libraries. -Contains the whole genome of an organism. -A restriction-enzyme is used to cut the genome (the DNA) at numerous locations. . polyA. cDNA. Reverse transcribe (and more). polyA.

cDNA Library
E N D
Presentation Transcript
cDNA Library MB206 Angelia 09
Making a DNA library Angelia 09
Types of Libraries Angelia 09 (3’UTR) (5’UTR)
Genomic DNA libraries -Contains the whole genome of an organism. -A restriction-enzyme is used to cut the genome (the DNA) at numerous locations. Angelia 09
polyA cDNA Reverse transcribe (and more) polyA polyA Clone in vector Digest DNA fragments Genomic Libraries cDNA library Genomic DNA mRNA Genomic DNA library Genomic DNA Angelia 09
polyA cDNA Reverse transcribe (and more) polyA polyA Clone in vector Digest DNA fragments cDNA Libraries cDNA library Genomic DNA mRNA Genomic DNA library Genomic DNA Angelia 09
“complementary” DNA [mRNA it is used to create it] cDNA Libraries • Purify mRNA • mRNA-> single stranded cDNA using reverse transcriptase. • Single stranded cDNA -> double stranded cDNA (DNA polymerase and other “cloning tricks”). • Linkers added to cDNA & clone into vectors as seen in genomic DNA libraries Angelia 09
Lodish, et al. 1999 Figure 7-14, 7-15 Angelia 09
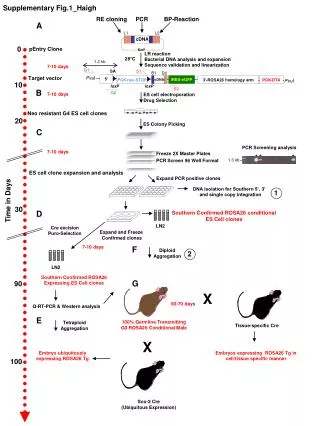
Genomic Libraries“The Details” Angelia 09
polyA Reverse transcribe (and more) cDNA polyA polyA Clone in vector Digest DNA fragments Genomic Libraries cDNA library Genomic DNA mRNA Genomic DNA library Genomic DNA Angelia 09
Digest genomic DNA with restriction enzymes • Which restriction enzyme should we select? Consideration: Try not to cut the gene or operon of interest • V. fischeri LUX operon 9kb Angelia 09
Selecting a restriction enzyme • What is the average size fragment that any given enzyme will yield? • If enzyme recognizes 6bp, statistically any given 6bp sequence will appear randomly every 46 bases (every 4096 bases) Angelia 09
How often does the enzymes cut? A ‘four-base cutter’ recognition sequence would occur once every 44 = 256 bp A ‘six-base cutter’ would give you fragments of about 4000 bp An ‘eight-base cutter’ recognition sequence would occur once every 48 or 65,536 bp. Angelia 09
How often do enzymes really cut? • But also we need to consider G+C content of the genome: • V. fischeri G+C content = 40% • Sal I recognizes sites that are G+C rich (GTCGAC) it will cut less often in Vibrio genomic DNA. Angelia 09
Let’s stop here and think about it! Angelia 09
Take your time! PROBLEM: • The genomic DNA is 60% G+C • You want a 5000bp fragment • Which enzyme would you try first, Why? • SmaI (CCCGGG) • EcoRI (GAATTC) • MseI (TTAA) • SacI (GAGCTC) Angelia 09
Digest DNA fragments Genomic Libraries Genomic DNA library Clone in vector Genomic DNA Angelia 09
Choosing a Vector • Usually you select a vector (plasmid, λ, other) depending on how big you want your DNA fragments to be & the capacity of the vector. Angelia 09
Common vectors used in library construction • Plasmids • Up to 10kb inserts • Modified Lambda phage • Up to 20kb inserts/40kbp for cosmids • Artificial cloning vectors • BAC- Vectors (bacterial artificial chromosome) Up to 100-150kbp inserts • YAC-Vectors (yeast artificial chromosome) Up to 500kbp inserts Angelia 09
Lambda Library Lodish, et al. Fig 7-12 Plasmid Library Lodish, et al. Fig 7-1 Angelia 09
Digest DNA fragments Genomic Libraries Genomic DNA library Clone in vector Genomic DNA Angelia 09
cDNA libraries • No cDNA library was made from prokaryotic mRNA. • Prokaryotic mRNA is very unstable • Genomic libraries of prokaryotes are easier to make and contain all the genome sequences. Angelia 09
cDNA libraries • cDNA libraries are very useful for eukaryotic gene analysis • Condensed protein encoded gene libraries, have much less junk sequences. • cDNAs have no introns genes can be expressed in E. coli directly • Are very useful to identify new genes • Tissue or cell type specific (differential expression of genes) Angelia 09
Most eukaryotic mRNAs are polyadenylated at their 3’ ends 5’ cap AAAAAAAAAAn • oligo (dT) can be bound to the poly(A) tail and used to recover the mRNA. mRNA isolation Angelia 09
Check the mRNA integrity Make sure that the mRNA is not degraded. Methods: Translating the mRNA : use cell-free translation system as wheat germ extract or rabbit reticulocyte lysate to see if the mRNAs can be translated Analysis the mRNAs by gel elctrophoresis: use agarose or polyacrylamide gels Angelia 09
Cloning the particular mRNAs Is useful especially one is trying to clone a particular gene rather to make a complete cDNA library. Fractionate on the gel: performed on the basis of size, mRNAs of the interested sizes are recovered from agarose gels Enrichment:carried out by hybridization Example: clone the hormone induced mRNAs (substrated cDNA library) Angelia 09
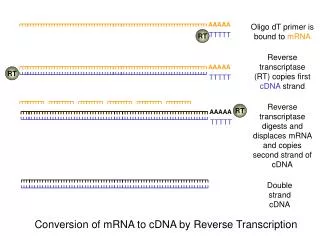
Synthesis of cDNA : First stand synthesis: materials as reverse transcriptase ,primer( oligo(dT) or hexanucleotides) and dNTPs (Fig 1.1) Second strand synthesis: best way of making full-length cDNA is to ‘tail’ the 3’-end of the first strand and then use a complementary primer to make the second. Angelia 09
mRNA 5’ AAAAA-3’ HO-TTTTTP-5’ Reverse transcriptase Four dNTPs mRNA 5’ AAAAA-3’ TTTTTP-5’ 3’ cDNA Terminal transferase dCTP mRNA 5’ AAAAA-3’ 3’-CCCCCCC TTTTTP-5’ cDNA Alkali (hydrolyaes RNA) Purify DNA oligo(dG) 5’-pGGGG-OH 3’-CCCCCCC TTTTTP-5’ cDNA Klenow polymerase or reverse Transcriotase Four dNTPs 5’-pGGGG -3’ 3’-CCCCCCC TTTTTP-5’ Duplex cDNA Angelia 09
Duplex cDNA 5’-pGGGG -3’ 3’-CCCCCCC TTTTTp-5’ Single strand-specific nuclease 5’-pGGGG -3’ 3’-CCC TTTTTp-5’ Klenow polymerase treat with E.coRI methylase 5’-pGGGG -3’ 3’-CCCC TTTTTp-5’ Add E.colRI linkers using T4 DNA ligase HO-CCG/AATTCGG-3’ 3’-GGCTTAA/GCC-OH HO-CCGAATTCGGGGGG CCGAATTCGG-3’ 3’-GGCTTAAGCCCCCC TTTTTGGCTTAAGCC-OH E.colRI digestion 5’-pAATTCGGGGGG CCG-3’ 3’-CCCCCCC TTTTTGGCTTAAp-5’ Ligate to vector and transfom Fig2.1 Second strand synthesis
Treatment of cDNA ends Blunt and ligation of large fragment is not efficient, so we have to use special acid linkers to create sticky ends for cloning. The process : Move protruding 3’-ends (strand-special nuclease) Fill in missing 3’ nucleotide(klenow fragment of DNA polyI and 4 dNTPs) Ligate the blunt-end and linkers(T4 DNA ligase) Tailing with terminal transferase or using adaptor molecules Restriction enzyme digestion(E.coRI )
Ligation to vector Any vectors with an EcoRI site would suitable for cloning the cDNA. The process : Dephosphorylate the vector with alkaline phosphatase Ligate vector and cDNA with T4 DNA ligase (plasmid or λ phage vector)
Screening The process of identifying one particular clone containing the gene of interest from among the very large number of others in the gene library . • Using nucleic acid probe to screen the library based on hybridization with nucleic acids. • Analyze the protein product. Angelia 09
Screening libraries Searching the genes of interest in a DNA library • Hybridization to identify the interested DNA or its RNA product • Radiolabeled probes which is complementary to a region of the interested gene • Probes: • An oligonucleotide derived from the sequence of a protein product of the gene • A DNA fragment/oligo from a related gene of another species • Blotting the DNA or RNA on a membrane • Hybridize the labeled probe with DNAmembrane (Southern) or RNA (Northern) membrane Angelia 09
Colony and plaque hybridization Transfer the DNA in the plaque or colony to a Nylon or nitrocellulose membrane Phage DNA bind to the membrane directly Bacterial colonies must be lysed to release DNA on the membrane surface. Hybridization (in a solution Containing Nucleic acid probe) (Alkali treatment) X-ray film(radio- actively labeled ) antibody or enzyme (modified nucleotide labeled Wash to remove unhybri- dization probe and visualize Line up the hybridizated region or repeated hybridization Angelia 09
Transfer to nitrocellulose or nylon membrane Keep master plate Select positive from master plate Denature DNA(NaOH) Bake onto membrane Probe with 32p-labled DNA complementary to gene of interest Expose to film Screening by plaque hybridization Angelia 09