cDNA SYNTHESIS
cDNA SYNTHESIS. Yaprak Dönmez December, 2009. RNA Ladder. RNA Ladder. W3 G2 F2 4 EN/DA F/IT. TH3 EC2 B/A 1 Y. RNA Ladder. 1.2% agarose, 70 V, 90 min. Determination of the RNA Concentration [RNA] μg/ml = A260 x dilution x 40.0 where


cDNA SYNTHESIS
E N D
Presentation Transcript
cDNA SYNTHESIS Yaprak Dönmez December, 2009
RNA Ladder RNA Ladder W3 G2 F2 4 EN/DA F/IT TH3 EC2 B/A 1 Y RNA Ladder 1.2% agarose, 70 V, 90 min
Determination of the RNA Concentration • [RNA] μg/ml = A260 x dilution x 40.0 • where • A260 = absorbance (in optical densities) at 260 nm • dilution = dilution factor (200) • 40.0 = average extinction coefficient of RNA.
cDNA Synthesis • Template: • Total RNA • Poly (A)+ RNA • Specific RNA
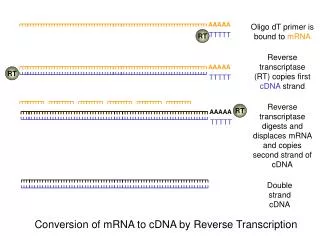
Priming • Oligo dT 18) • Adv:You can synthesize complete cDNAs, beginning at the poly A+ tail and ending at the 5 end of the mRNA. • Disadv: The reverse transcriptases used to synthesize cDNAs have an average length of 1 to 2 kb, whereas mRNAs can easily be 10 kb long. The protein coding portion of interest is usually in the vicinity of the 5 end!
Random hexamer • Hybridizes somewhere along the mRNA so that all mRNA segments are represented in the cDNA: • Gene specific primers • Adv: specific mRNA • Disadv: lower yields
Reverse Transcriptases (RTs) • The reverse transcriptase from the avian myoblastosis virus (AMV-RT): • Temperature optimum of activity at 45-50°C. • Highly thermostable, can be used at temperatures up to 60°C. • Demonstrates DNA exonuclease and RNase activities. • The reverse transcriptase from the Moloney murine leukemia virus (MMLV-RT): • The optimal working temperature is about 37C, with a maximum of 42C. • Weaker RNase activity.
Protocol for First-strand cDNA Synthesis Mix and briefly centrifuge all components after thawing, keep on ice. DEPC treated dH2O to 11 µL
Heat in 70° for 5 min, chill for a few minutes. This will disrupt secondary structures formed in the RNA to make it rather linear for the primer to pair to it. • Add:
Heat in 37° for 5 min, chill for a few minutes. This will anneal primer to complementary RNA. • Add 0.3 µL M-MuLV Reverse Transcriptase. • Incubate at 42° for 60 min. Extension of primer will occur via RT. • Heat-inactivate reverse transcriptase at 72° for 10 min.
Normalization • Most gene expression assays are based on the comparison of two or more samples and require uniform sampling conditions for this comparison to be valid. • Many factors can contribute to variability in the analysis of samples, making the results difficult to reproduce between experiments: • Sample degradation, extraction efficiency, contamination → RNA isolation • Sample concentration, RNA integrity, the reagents used, presence of contaminants → reverse transcription • Housekeeping genes such as ß-actin, ß-tubulin, GAPDH, and18S ribosomal RNA have often been used as reference genesfor normalization, with the assumption that the expression ofthese genes is constitutively high and that a given treatment willhave no effect on the expression level.
References: • http://hominid.uchicago.edu/ProtocolPDFs/OligodTcDNASynthesis.pdf • http://www.ncbi.nlm.nih.gov/books/bv.fcgi?highlight=cDNA,Synthesis&rid=mcb.section.1611#1619 • http://www.fermentas.com/catalog/modifyingenzymes/m_mulvrt.html • Kok J B et.al. Normalization of gene expression measurements in tumor tissues: comparison of 13 endogenous control genes. Laboratory Investigation 2005; 85, 154–159. • http://www.dna.ohiou.edu/literature/qRT_pCR/Normalization_Methods_for_qPCR.pdfl • Farrell R E (2005). RNA Methodologies ISBN 0122496965, 9780122496967.