Expression of clones genes
260 likes | 540 Vues
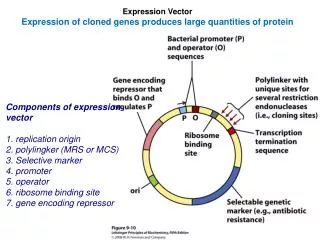
Expression of clones genes. a.o. Primrose & Twyman, 7th edition, pp.81-93 Primrose, Twyman & Old, 6th edition, pp.70-83. IPTG-induced expression of RP4 proteins : (A) DNA primase (118 and 80 kDa proteins) ; (B) 16.5 and

Expression of clones genes
E N D
Presentation Transcript
Expression of clones genes a.o. Primrose & Twyman, 7th edition, pp.81-93 Primrose, Twyman & Old, 6th edition, pp.70-83
IPTG-induced expression of RP4 proteins : (A) DNA primase (118 and 80 kDa proteins) ; (B) 16.5 and 8.6 kDA proteins. Host strain : E. coli HB101. Analysis by SDS-polyacrylamide gel electrophoresis. (A) insert in direct (c, d) or opposite (e,f) orientation ; induced (c, e) or non-induced (d,f) with IPTG. (a) reference proteins ; (b) purified RP4 DNA primase (B) insert in direct (b, c) or opposite (d,e) orientation ; induced (b, d) or non-induced (c,e) with IPTG. (f) reference proteins ; (a) purified 16.5 kDa protein.
E. coli Pribnow box (-10) T89 A99 T50 A65 A65 T100 s70 binding site (-35) T85 T83 G81 A61 C69 A52 E. coli promoters, and hybrid promoters tacI and II E. coli transcriptional termination (r-independent)
Codeword usage in strongly and weakly expressed E. coli genes. "strong" : 24 genes (5253 triplets), a.o. 12 of ribosomal proteins, 7 of outer membrane proteins, and some for RNA polymerase subunits and translation initiation and termination factors. "weak" : 18 genes (5231 triplets), a.o. for different repressor proteins, lac-permease, etc.
Control of expression by the l CI repressor on the Ptrp promoter
Expression by the PBAD promoter and its regulation (in pBAD vectors).
Inclusion bodies in E. coli (Scanning electron micrograph) (Thin section through the cells)
Constitutive expression of the cat gene, and fusions by random cloning into the ScaI site (lane 2) (lanes 1, 6, 7, 9-15) Lanes 4 and 5 are plasmid-free cells ; lanes 3 and 8 are protein size markers.
Different types of translational fusion constructs. S : signal sequence ; I : membrane integration domain ; A, B : fusion peptides, e.g. affinity tags
Protein production and processing with the pBAD/His vectors C-terminal fusion 6 HIS-residues for affinity purification cleavage by enterokinase (D-D-D-D-K-*) after processing a few extra amino acids remain at the N-terminus.
C-terminal fusion at biotin carboxylase Binding onto streptavidin column for purification. Processing with factor Xa protease (I-E-G-R-* of I-D-G-R-*(not followed by P or R))
~100 amino acids ~50 amino acids Splicing of an intein
Processing of N-terminal fusions at an intein, immobilized onto chitin columns
pMAL expression vectors malE : encodes maltose binding protein.
Properties of pMAL vectors : • ColE1 ori ; M13 ori ; Ptac ; rrnB terminators ; lacIq ; bla • Fusion construct : malE - DDDDDDDDDD - IEGR - MCS(polylinker) - lacZa • pMAL-c2 : malE signal peptide sequence deleted • pMAL-p2 : malE signal peptide sequence : secretion to periplasm • fusion in XmnI : exact joining of target protein at factor Xa sequence • 10 Asp residues (D) separate the two fusion moieties • insertion in EcoRI is identical (in reading frame) as in lgt11 (lacZ) Cleavage site offactor Xa I - E - G - R * XmnI nnn nnn nGA Ann nnT TCn nnn ATC-GAG-GGA-AGG-ATT-TCA-GAA-TTC- EcoRI
Multiple cloning site in (one of) the pMAL vectors. Other possibility (in some vectors) Pro-Gly-Ala-Ala-His-Tyr : cleavage by Genenase I = engineered form of subtilisin : cleaves beyond His-Tyr (depending on the next residue) (cleaves His-Tyr-Glu and His-Tyr-Asp slowly, doesn't cleave His-Tyr-Pro or His-Tyr-Ile)
Processing of expression products from fusion constructs in pMAL vectors.
(Commercial) Pinpoint vectors two promoters : T7 and tac factor Xa cleavage site N-terminal biotinylation (tag) polylinker in vitro RNA probes by SP6 promoter secretion signal (some vectors) Detection by streptavidin-alkaline phosphatase. Elution of fusion protein with 5 mM biotin. (hence : no denaturation required) Biotinylation at a Lys residue(a single biotin)in E. coli by the biotin ligase holoenzyme. Accessible to avidin or streptavidin ; tag for detection and purification. nb. E. coli produces a single endogenous biotinylated protein that, in its native conformation, does not bind to avidin : => highly specific for the recombinant fusion protein