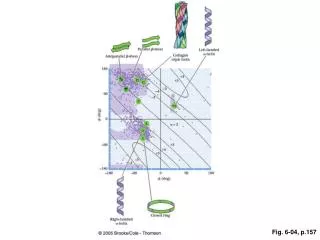
Fig. 6-7
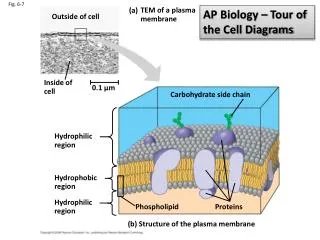
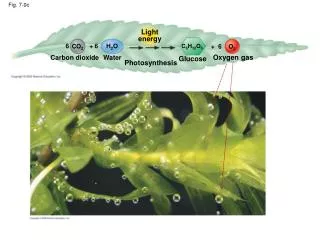
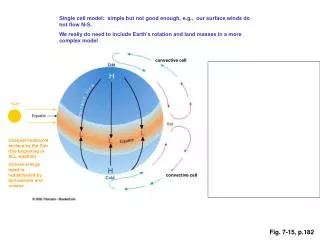
Fig. 6-7. AP Biology – Tour of the Cell Diagrams . (a). TEM of a plasma membrane. Outside of cell. Inside of cell. 0.1 µm. Carbohydrate side chain. Hydrophilic region. Hydrophobic region. Hydrophilic region. Phospholipid. Proteins. (b) Structure of the plasma membrane.

Fig. 6-7
E N D
Presentation Transcript
Fig. 6-7 AP Biology – Tour of the Cell Diagrams (a) TEM of a plasma membrane Outside of cell Inside of cell 0.1 µm Carbohydrate side chain Hydrophilic region Hydrophobic region Hydrophilic region Phospholipid Proteins (b) Structure of the plasma membrane
Fig. 6-8 Surface area increases while total volume remains constant 5 1 1 Total surface area [Sum of the surface areas (height width) of all boxes sides number of boxes] 150 750 6 Total volume [height width length number of boxes] 1 125 125 Surface-to-volume (S-to-V) ratio [surface area ÷ volume] 6 1.2 6
Fig. 6-9a Nuclear envelope ENDOPLASMIC RETICULUM (ER) NUCLEUS Nucleolus Rough ER Smooth ER Flagellum Chromatin Centrosome Plasma membrane CYTOSKELETON: Microfilaments Intermediate filaments Microtubules Ribosomes Microvilli Golgi apparatus Peroxisome Mitochondrion Lysosome
Fig. 6-9b Rough endoplasmic reticulum Nuclear envelope Nucleolus NUCLEUS Chromatin Smooth endoplasmic reticulum Ribosomes Central vacuole Golgi apparatus Microfilaments Intermediate filaments CYTO- SKELETON Microtubules Mitochondrion Peroxisome Chloroplast Plasma membrane Cell wall Plasmodesmata Wall of adjacent cell
Fig. 6-10 Nucleus 1 µm Nucleolus Chromatin Nuclear envelope: Inner membrane Outer membrane Nuclear pore Pore complex Rough ER Surface of nuclear envelope Ribosome 1 µm 0.25 µm Close-up of nuclear envelope Pore complexes (TEM) Nuclear lamina (TEM)
Fig. 6-11 Cytosol Endoplasmic reticulum (ER) Free ribosomes Bound ribosomes Large subunit Small subunit 0.5 µm Diagram of a ribosome TEM showing ER and ribosomes
Fig. 6-12 Smooth ER Nuclear envelope Rough ER ER lumen Cisternae Transitional ER Ribosomes Transport vesicle 200 nm Rough ER Smooth ER
Fig. 6-13 cis face (“receiving” side of Golgi apparatus) 0.1 µm Cisternae trans face (“shipping” side of Golgi apparatus) TEM of Golgi apparatus
Fig. 6-14 1 µm Nucleus Vesicle containing two damaged organelles 1 µm Mitochondrion fragment Peroxisome fragment Lysosome Digestive enzymes Lysosome Lysosome Plasma membrane Peroxisome Digestion Food vacuole Digestion Mitochondrion Vesicle (a) Phagocytosis (b) Autophagy
Fig. 6-17 Intermembrane space Outer membrane Free ribosomes in the mitochondrial matrix Inner membrane Cristae Matrix 0.1 µm
Fig. 6-18 Ribosomes Stroma Inner and outer membranes Granum 1 µm Thylakoid
Fig. 6-21 Vesicle ATP Receptor for motor protein Motor protein (ATP powered) Microtubule of cytoskeleton (a) Microtubule Vesicles 0.25 µm (b)
Table 6-1 10 µm 10 µm 10 µm Column of tubulin dimers Keratin proteins Actin subunit Fibrous subunit (keratins coiled together) 25 nm 8–12 nm 7 nm Tubulin dimer
Fig. 6-22 Centrosome Microtubule Centrioles 0.25 µm Microtubules Longitudinal section of one centriole Cross section of the other centriole
Fig. 6-24 Outer microtubule doublet Plasma membrane 0.1 µm Dynein proteins Central microtubule Radial spoke Protein cross-linking outer doublets Microtubules (b) Cross section of cilium Plasma membrane Basal body 0.5 µm 0.1 µm (a) Longitudinal section of cilium Triplet (c) Cross section of basal body
Fig. 6-25 Microtubule doublets ATP Dynein protein (a) Effect of unrestrained dynein movement ATP Cross-linking proteins inside outer doublets Anchorage in cell (b) Effect of cross-linking proteins 1 3 2 (c) Wavelike motion
Fig, 6-27a Muscle cell Actin filament Myosin filament Myosin arm (a) Myosin motors in muscle cell contraction
Fig. 6-28 Secondary cell wall Primary cell wall Middle lamella 1 µm Central vacuole Cytosol Plasma membrane Plant cell walls Plasmodesmata
Fig. 6-30 Polysaccharide molecule Proteoglycan complex Collagen EXTRACELLULAR FLUID Carbo- hydrates Fibronectin Core protein Integrins Proteoglycan molecule Plasma membrane Proteoglycan complex CYTOPLASM Micro- filaments
Fig. 6-32 Tight junction Tight junctions prevent fluid from moving across a layer of cells 0.5 µm Tight junction Intermediate filaments Desmosome Desmosome Gap junctions 1 µm Extracellular matrix Space between cells Gap junction Plasma membranes of adjacent cells 0.1 µm