Methods
Methods. Step I – FLUX Analysis. MILP => LP: Setting new flux bounds for highly/lowly expressed genes (=> reactions) ? sampling ?. Methods. * Step II – Reaction Profiling over NCI-60. Reaction Flux Similarity. * Step II – Sequence information.

Methods
E N D
Presentation Transcript
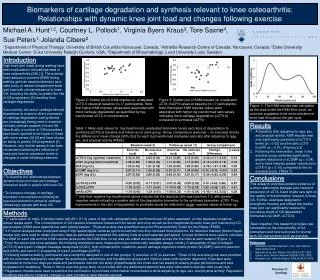
Methods Step I – FLUX Analysis • MILP => LP: • Setting new flux bounds for highly/lowly expressed genes (=> reactions) ? • sampling ?
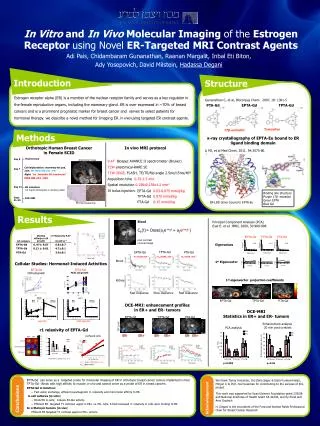
Methods * Step II – Reaction Profiling over NCI-60 Reaction Flux Similarity * Step II – Sequence information S_TS (s,t) Sequence similarity Smith-Waterman Combined Measure
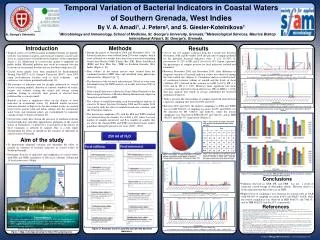
Methods * Step III – ML, based on Drug - Reaction Network
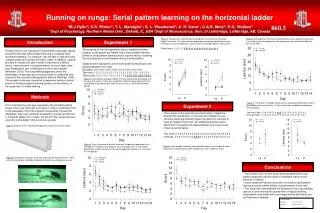
Results * Building the Drug-Reaction Network (DRN) * Reaction interact with a drug through the same enzyme (dashed) or not (Solid)
Drugs: 339/1492 50% Clear clusters: * CNS * Antineoplastic + Blood + Cardiovascular + respiratory Reactions: 811/2299 * CNS <=> Transport * Cancer <=> Nucleotides 145
* Reaction Profiling (flux similarity) Only about 350 reaction which are active in at least 1 cell line Both Reaction and Cell lines show 2 principle groups
R1 E D R2 R1 E1 D R2 E2
* Predictions Kernel KNN with sequence similarity
Conclusion metabolic flux similarity-measures provide good tools for drug-target prediction Combining metabolic flux and sequences similarities-measures improves performance Elaborate on the metabolic mechanisms involved with drug application