ENZYMES
ENZYMES. History of Enzymes. -1700s and early 1800s, the digestion of meat by stomach secretions and the conversion of starch to sugars by plant extracts and saliva were known. --mechanism by which this occurred had not been identified. History of Enzymes.

ENZYMES
E N D
Presentation Transcript
History of Enzymes -1700s and early 1800s, the digestion of meat by stomach secretionsand the conversion of starch to sugars by plant extracts and saliva were known. --mechanism by which this occurred had not been identified.
History of Enzymes -19th century, when studying the fermentation of sugar to alcohol by yeast, Louis Pasteur came to the conclusion that it was catalyzed by a vital force contained within the yeast cells called "ferments", which were thought to function only within living organisms. --He wrote that "alcoholic fermentation is an act correlated with the life and organization of the yeast cells, not with the death or putrefaction of the cells. Yeast
History of Enzymes • 1878: German physiologist Wilhelm Kühne • first used the term enzyme, which literally means “in yeast” • 1897: Eduard Buchner • began to study the ability of yeast extracts that lacked any living yeast cells to ferment sugar. He also found that the sugar was fermented even when there were no living yeast cells in the mixture. • He named the enzyme that brought about the fermentation of sucrose "zymase". In 1907 he received the Nobel Prize in Chemistry “for his biochemical research and his discovery of cell-free fermentation".
Enzymes • Enzymes are biomolecules that catalyze, increase the rates of chemical reactions without being altered during the reaction. • Almost all enzymes are proteins; • Enzymes are essential to life. • In enzymatic reactions, the molecules at the beginning of the process are called substrates, and the enzyme converts them into different molecules, the products.
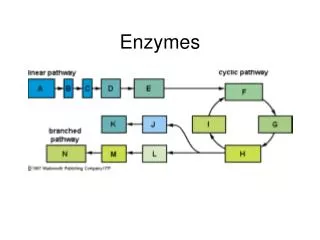
Enzymes • Almost all processes in a biological cell need enzymes in order to occur at significant rates. • Since enzymes are extremely selective for their substrates and speed up only a few reactions from among many possibilities, the set of enzymes made in a cell determines which metabolic pathways occur in that cell.
Enzymes • Enzyme activity can be affected by other molecules. • Inhibitors are molecules that decrease enzyme activity. • Inducers are molecules that increase activity. Many drugs and poisons are enzyme inhibitors. • Activity is also affected by temperature, chemical environment (e.g. pH),
ENZYMES are biological catalyst ENZYME CHARACTERISTICS • The basic function of an enzyme is to increase the rate of a reaction • Most enzymes act specifically with only one reactant (called a substrate) to produce products • The most remarkable characteristic is that enzymes are regulated from a state of low activity to high activity and vice versa
Three-dimensional structure of an ENZYME • Enzymes are proteins • They have a globular shape • A complex 3-D structure Human pancreatic amylase
Enzyme Structure • Most enzymes are proteins • Enzymes may require a non-peptide component as a cofactor. The peptide component is called the apoenzyme, the cofactor is called as the coenzyme and the combined functional unit is the holoenzyme • Cofactors that are tightly bound to the polypeptide are called prosthetic groups. Such proteins are called as complex or conjugated proteins. Proteins without prosthetic groups are simple proteins
The Active Site • One part of an enzyme, the active site, is particularly important • The shape and the chemical environment inside the active site permits a chemical reaction to proceed more easily
APOENZYME • May be inactive in its original synthesized structure PROENZYME OR ZYMOGEN • The inactive form of the apoenzyme • May contain several extra amino acids in the protein which are removed, and allows the final specific tertiary structure to be formed before it is activated as an apoenzyme
The Substrate • The substrate of an enzyme are the reactants that are activated by the enzyme; • Enzymes are specific to their substrates; • The specificity is determined by the active site.
COFACTOR • An additional non-protein molecule that is needed by some enzymes to help the reaction Nitrogenase enzyme with Fe, Mo and ADP cofactors
COFACTOR • A non-protein substance which may be organic and called coenzyme • Common coenzymes are vitamins and metal ions
COFACTOR • Another type of cofactor is an inorganic metal ion called a metal ion activator • Are inorganic and may be bonded through coordinate covalent bonds • Metal ions as Zn+2, Mg+2, Mn+2, Fe+2, Cu+2, K+, and Na+1 are used in enzymes as cofactors
PROSTHETIC GROUPS • Are tightly incorporated into protein structure by covalent or noncovalent forces • Examples include derivatives of B vitamins such as pyridoxal phosphate, flavin mononucleotide (FMN), flavin adenine dinucleotide (FAD), thiamin pyrophosphate, biotin and METAL IONS of Co, Cu, Mg, Mn, and Zn. • METALLOENZYMES – enzymes that contain tightly bound metal ions
NOMENCLATURE • The commonly used names for most enzymes describe the type of reaction catalyzed, followed by the suffix –ase. • Dehydrogenases – remove hydrogen atoms • Proteases – hydrolyze proteins • Isomerases – catalyze rearrangement in configuration
NOMENCLATURE • Modifiers may precede the name to indicate; (a) the substrate (xanthineoxidase) (b) the source of the enzyme (pancreaticribonuclease) (c) its regulation (hormone-sensitive lipase) (d) a feature of its mechanism of action (cysteine protease)
NOMENCLATURE • Alphanumeric designators may be added to identify multiple forms of an enzyme ( eg., RNA polymerase III; protein kinase C ) • Some enzymes retain their original trivial names, which give no hint of the associated enzymatic reaction • Examples are pepsin, trypsin, and chymotrypsin which catalyzes the hydrolysis of proteins
Classification of Enzymes Based on catalyzed reactions, the nomenclature committee of the International Union of Biochemistry and Molecular Biology (IUBMB) recommended the following classification: • OXIDOREDUCTASES • Catalyze a variety of oxidation-reduction reactions • Common names include dehydrogenase, oxidase, reductase and catalase
Classification of Enzymes 2. TRANSFERASES • Catalyze transfers of groups (acetyl, methyl, phosphate, etc.). • The first three subclasses play major roles in the regulation of cellular processes. • The polymerase is essential for the synthesis of DNA and RNA.
Three major regulatory chemical reactions. (a)Acetylation - addition of an acetyl group to lysine's R group by acetyltransferase. (b)Methylation - addition of a methyl group to DNA's base (e.g. cytosine) by methylase. (c)Phosphorylation - addition of a phosphate group to the R group of tyrosine, serine or threonine (only tyrosine is shown here) by protein kinase.
Classification of Enzymes 3. HYDROLASES Catalyze hydrolysis reactions where a molecule is split into two or more smaller molecules by the addition of water • PROTEASES split protein molecules • HIV protease is essential for HIV replication • Caspase plays a major role in apoptosis • NUCLEASES split nucleic acids (DNA and RNA) • Based on the substrate type, they are divided into RNase and DNase. • RNase catalyzes the hydrolysis of RNA • DNase acts on DNA
Classification of Enzymes • Nucleases cont… • They may also be divided into exonuclease and endonuclease. • The exonuclease progressively splits off single nucleotides from one end of DNA or RNA. • The endonuclease splits DNA or RNA at internal sites. • PHOSPHATASE catalyzes dephosphorylation (removal of phosphate groups). • Example: calcineurin (also known as protein phosphatase 3) • The immunosuppressive drugs Tacrolimus, Sirolimus, Everolimus and Cyclosporin A are the calcineurin inhibitors
Classification of Enzymes 5. ISOMERASES • Catalyze atomic rearrangements within a molecule. • Examples include rotamase, protein disulfide isomerase (PDI), epimerase and racemase 4. LYASES • Catalyze the cleavage of C-C, C-O, C-S and C-N bonds by means other than hydrolysis or oxidation. • Common names include decarboxylase and aldolase.
The role of rotamase and protein disulfide isomerase (PDI). The reactions catalyzed by the two enzymes can assist a peptide chain to fold into a correct three-dimensional structure
6. LIGASES • Catalyze the reaction which joins two molecules • Examples include peptide synthase, aminoacyl-tRNA synthetase, DNA ligase and RNA ligase The IUBMB committee also defines subclasses and sub-subclasses • Each enzyme is assigned an EC (Enzyme Commission) number • For example, the EC number of catalase is EC1.11.1.6 • The first digit indicates that the enzyme belongs to oxidoreductase (class 1) • Subsequent digits represent subclasses (1.11. acting on a peroxide as acceptor) and sub-subclasses (1.11.1peroxidases)
Mechanism of Enzyme Action The molecule acted upon a unique geometric shape that is complementary to the geometric shape of a substrate molecule
Mechanism of Enzyme Action Lock and Key Theory • first postulated in 1894 by Emil Fischer • The lock is the enzyme and the key is the substrate • Only the correctly sized key (substrate) fits into the key hole (active site) of the lock (enzyme)
S E E E Enzyme may be used again Enzyme-substrate complex P P Reaction coordinate The Lock and Key Hypothesis
Postulated by Daniel Koshland It states that, when substrates approach and bind to an enzyme they induce a conformational change This change is analogous to placing a hand (substrate) into a glove (enzyme) The Induced Fit Theory
The Induced Fit Theory • Some proteins can change their shape (conformation) • When a substrate combines with an enzyme, it induces a change in the enzyme’s conformation • The active site is then moulded into a precise conformation • Making the chemical environment suitable for the reaction • The bonds of the substrate are stretched to make the reaction easier (lowers activation energy)
Hexokinase (a) without (b) with glucose substrate http://www.biochem.arizona.edu/classes/bioc462/462a/NOTES/ENZYMES/enzyme_mechanism.html The Induced Fit Theory • This explains the enzymes that can react with a range of substrates of similar types
The Induced Fit Theory • Assumes that the substrate plays a role in determining the final shape of the enzyme and that the enzyme is partially flexible. • This explains why certain compounds can bind to the enzyme but do not react because the enzyme has been distorted too much • Other molecules may be too small to induce the proper alignment and therefore cannot react • Only the proper substrate is capable of inducing the proper alignment of the active site
This is a representation of carboxypeptidase A with a substrate (turquoise) bound in the active site. The active site is in the induced conformation. This is a molecular model of the unbound carboxypeptidase A enzyme
MECHANISMS TO FACILITATE CATALYSIS A. CATALYSIS BY PROXIMITY • For molecules to react, they must come within bond-forming distance of one another • The higher the concentration, the more frequently they will encounter one another and the greater will be their rate of interaction • aka entropy reduction • ACID-BASE CATALYSIS • Can be specific or general • “Specific” meaning only protons (H3O+ , specific acid) or OH- ions (specific base)
MECHANISMS TO FACILITATE CATALYSIS • Proximity: Reaction between bound molecules doesn't require an improbable collision of 2 molecules -- they're already in "contact" (increases the local concentration of reactants). • Orientation: Reactants are not only near each other on enzyme, they're oriented in optimal position to react, so the improbability of colliding in correct orientation is taken care of.
Rate enhancement by entropy reduction. a) bimolecular reaction(high activation energy, low rate)
Rate enhancement by entropy reduction. • b) unimolecular reaction, rate enhanced by factor of 105 due to increased probability of collision/reaction of the 2 groups.
Rate enhancement by entropy reduction. • c) constraint of structure to orient groups better (elimination of freedom of rotation around bonds between reactive groups), rate enhanced by another factor of 103, for 108 total rate enhancement over bimolecular reaction.