References
On Computing Greyscale Morphology with Large Exact Spheres in Arbitrary Dimensions Via 1-D Distance Transforms. Mathematics, Informatics and Statistics.

References
E N D
Presentation Transcript
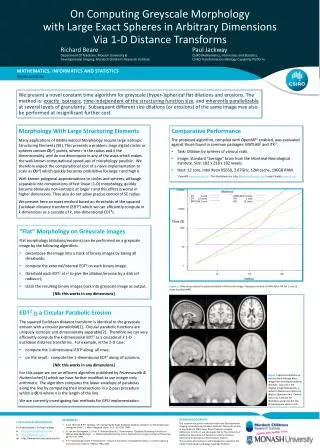
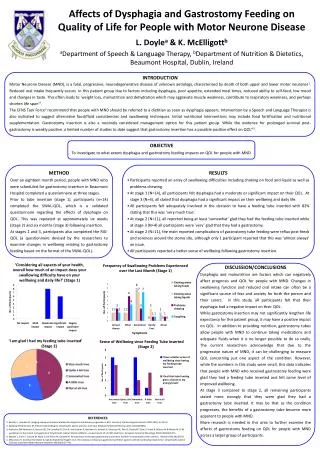
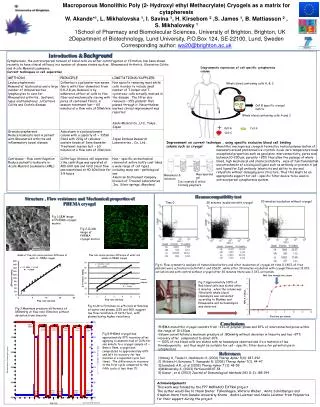
On Computing Greyscale Morphology with Large Exact Spheres in Arbitrary Dimensions Via 1-D Distance Transforms Mathematics, Informatics and Statistics We present a novel constant time algorithm for greyscale (hyper-)spherical flat dilations and erosions. The method is: exactly isotropic, time-independent of the structuring function size, and inherently parallelizable at several levels of granularity. Subsequent different size dilations (or erosions) of the same image may also be performed at insignificant further cost. Richard Beare Paul Jackway Department Of Medicine, Monash University & CSIRO Mathematics, Informatics and Statistics, Developmental Imaging, Murdoch Children’s Research Institute CSIRO Transformational Biology Capability Platform Dr Richard Beare / Dr Paul Jackway e Richard.Beare@monash.edu e Paul.Jackway@csiro.au w http://www.mcri.edu.au/research/ Morphology With Large Structuring Elements Many applications of Mathematical Morphology require large isotropic Structuring Elements (SE). This presents a problem: large digital circles or spheres contain O(rk) points, where r is the radius and k the dimensionality, and do not decompose in any of the ways which makes the well-known computational speed-ups of morphology possible. We therefore expect the computational cost of a naive implementation to scale as O(rk) which quickly becomes prohibitive for large r and high k. Well-known polygonal approximations to circles and spheres, although separable into compositions of fast linear (1-D) morphology, quickly become obviously non-isotropic at larger r and this effect is worse in higher dimensions. They also do not allow precise control of SE radius. We present here an exact method based on thresholds of the squared Euclidean distance transform (EDT2) which we can efficiently compute in k-dimensions as a cascade of k, one-dimensional EDT2s. Comparative Performance The proposed algorithm, compiled with OpenMP1 enabled, was evaluated against those found in common packages: MATLAB2and ITK3. Task: Dilation by spheres of various radii. Image: Standard “average” brain from the Montreal Neurological Institute. Size: 182 x 218 x 182 voxels. Host: 12 core, Intel Xeon X5650, 2.67GHz, 12M cache, 196GB RAM. 1 OpenMP www.openmp.org2 The MathWorksInc. http://www.mathworks.com3 Insight Toolkit www.itk.org DT 1core DT 12 cores MATLAB 1 core MATLAB 12 cores ITK 1 core ITK 12 cores Time (S) “Flat” Morphology on Greyscale Images Flat morphology (dilations/erosions) can be performed on a greyscale image by the following algorithm: • decomposethe image into a stack of binary images by taking all thresholds; • computethe external/internal EDT2 on each binary image; • thresholdeach EDT2 at r2 to give the dilation/erosion by a disk (of radius=r); • stack the resulting binary images back into greyscale image as output. [NB: this works in any dimensions] Figure 1:Time versus radius for spherical dilation of the brain image. Proposed method vs MATLAB vs ITK for 1 and 12 cores (via OpenMP). EDT2is a Circular Parabolic Erosion The squared Euclidean distance transform is identical to the greyscale erosion with a circular paraboloid[1]. Circular parabolic functions are uniquely isotropic and dimensionally separable[2]. Therefore we can very efficiently compute the k-dimensional EDT2 as a cascade of k 1-D Euclidean distance transforms. For example, in the 2-D case: • computethe 1-dimensional EDT2 along all rows; • on the result: compute the 1-dimensional EDT2 along all columns. [NB: this works in any dimensions] For this paper we use an efficient algorithm published by Felzenszwalb & Huttenlocher[3] which we have further modified to use integer only arithmetic. The algorithm computes the lower envelope of parabolas along the line by computing their intersections in a 2-pass procedure which is O(n) where n is the length of the line. We are currently investigating fast methods for GPU implementation. Figure 2 Spherical dilations of the standard average brain image from the Montreal Brain Institute. Top row is the original image followed by a radius 5 dilation and radius 10 dilation (bottom row). Central slice only is shown for illustrative purposes but the processing was done in 3-D. Acknowledgements This research was partly conducted within the Developmental Imaging research group, Murdoch Children’s Research Institute at the Children’s MRI Centre, Royal Children's Hospital, Melbourne Victoria. It was also supported by The University of Melbourne Department of Paediatrics and the Victorian Government's Operational Infrastructure Support. The second author wishes to acknowledge the support of the CSIRO Transformational Biology Capability Platform. • References • A.J.H. Mehnert & P.T. Jackway. “On Computing the Exact Euclidean Distance Transform on Rectangular and Hexagonal Grids”, J. Math. Imaging & Vision, 11(3): 223-230, 1999. • R. van den Boomgaard, L. Dorst, S. Makram-Ebeid & J. Schavemaker, "Quadratic Structuring Functions in Mathematical Morphology", in: Mathematical Morphology and its Applications to Image and Signal Processing, pp 147-154, 1996. • P. F. Felzenszwalb and D. P. Huttenlocher. "Distance Transforms of Sampled Functions", Cornell Computing and Information Science, TR2004-1963, 2004.