Protein Synthesis
240 likes | 347 Vues
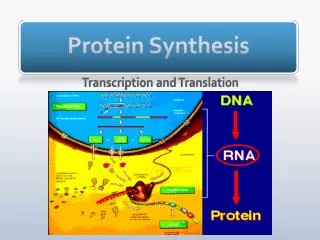
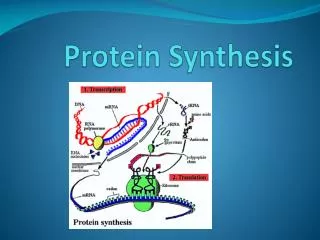
Explore the processes of transcription, mRNA processing, translation, and more in protein synthesis. Learn about ribosomes, tRNA, genetic codes, and the differences between prokaryotic and eukaryotic systems.

Protein Synthesis
E N D
Presentation Transcript
Transcription: Initiation • RNA polymerase attaches to a “promoter” region in front (“upstream”) of a gene • Eukaryotes: RNA polymerase requires multiple “Transcription factor” proteins to be able to bind to the promoter • Promoters have characteristic DNA sequences (ex:TATA Box in eukaryotes – 5’ TATAAA 3’)
Transcription: Elongation • RNA production occurs in a 5’ to 3’ direction • “Template Strand” of DNA is the one that the RNA transcript is produced off of • The “Coding Strand” of the DNA will have the same sequence as the RNA transcript…EXCEPT?
Transcription: Termination • mRNA transcript production continues until the end of the transcription unit is reached • Multiple mechanisms of termination exist
Post-Transcriptional mRNA processing (EUKARYOTES ONLY!) • Exons are the “coding” regions of the DNA and Introns are the “noncoding” regions of the DNA
How is the mRNA transcript modified? • The introns must be removed and the exons must be spliced together prior to the mRNA leaving the nucleus • **This is required to produce a functional protein
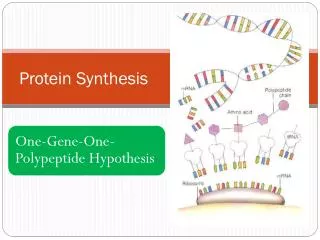
How does alternative splicing contribute to the production of various proteins? • Having multiple exons in a gene allows eukaryotes to make multiple functioning proteins from one gene
Ribosomes • Site of Protein Synthesis • “non-membrane” bound organelle • All cells have them • Composed of two subunits • Has 3 sites: • A site – where amino acids enter the ribosome • P site – where the growing polypeptide chain is • E site – where empty tRNA molecules leave
tRNA • Transfer RNA molecules • brings amino acids to the ribosome • Amino-acyl synthase enzyme catalyzes reaction to attach amino acid to tRNA • When carrying amino acid, “charged tRNA” • Anticodon region interacts with mRNA
The Genetic Code • Universal across ALL domains of Life! • Triplet Code: mRNA is read in units of three bases (codons) • There are 64 possible codons (for 20 amino acids) • The code is redundant and unambiguous • The code has “start” and “stop” codons
Translation: Initiation • mRNA attaches to the small ribosomal subunit • Methionine is brought to the start codon (AUG) by the methionine tRNA • Start Codon is located in the P-site • tRNA binding is mediated by its “anti-codon” • “Translation Initiation Complex”
Translation: Elongation • The next codon determines the next amino acid to be brought to the ribosome • Incoming, “Charged” tRNA enters at the A-site • Growing polypeptide chain is transferred to the new tRNA molecule – Peptide Bond Formed • Ribosome translocates (shifts) tRNA molecule with chain to P-site; previous (now “uncharged”) tRNA moves to E-site • Next codon available in A-site for new t-RNA
Translation: Termination • When a stop codon (UAG, UAA, or UGA) is encountered, a release factor binds to the A-site • The polypeptide chain is released • The ribosome disassembles
How does prokaryotic protein synthesis compare to eukaryotic?
How does prokaryotic compare to eukaryotic protein synthesis? • Transcription: Prokaryotes can bind DIRECTLY to the promoter region (no transcription factors required) • No Post-transcription modification of RNA • Prokaryotes have no nucleus so Transcription and Translation are coupled (occur together) – RNA is transcribed and translated at the same time
What is a polyribosome? How do prokaryotic and eukaryotic organisms benefit from them? • Many ribosomes translating the same RNA transcript • Enables simultaneous translation of one transcript