Molecular Biology
570 likes | 806 Vues
Molecular Biology. Part I: Chemistry and Genetics Part II: Maintenance of the Genome Part III: Expression of the Genome Part IV: Regulation Part V: Methods. Ch 6: The structures of DNA and RNA Ch 7: Chromosomes, chromatins and the nucleosome Ch 8: The replication of DNA

Molecular Biology
E N D
Presentation Transcript
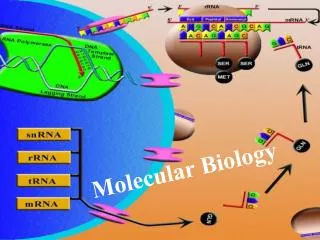
Molecular Biology Part I: Chemistry and Genetics Part II: Maintenance of the Genome Part III: Expression of the Genome Part IV: Regulation Part V: Methods
Ch 6: The structures of DNA and RNA Ch 7: Chromosomes, chromatins and the nucleosome Ch 8: The replication of DNA Ch 9: The mutability and repair of DNA Ch 10: Homologous recombination at the molecular level Ch 11: Site-specific recombination and transposition of DNA
Consider the structure of DNA within the cell, and the biological relevance of the structure. • DNA is associated with proteins in cells, both prokaryotes and eukaryotes, even viruses • Each DNA and its associated proteins is called a chromosome
Vocabulary Nucleus: 细胞核; Nucleolus: 核仁 Nucleoid: 类核 Mitosis: 有丝分裂;Meiosis:减数分裂 interphase:分裂间期 Histone: 组蛋白;Nucleosome: 核小体 Chromotasome: 染色小体 Chromosome: 染色体; Chromatin: 染色质;eu-; hetero- Centromere (中心粒) Telomere(端粒) Repetitive DNA (重复DNA) Tandem gene cluster(串联基因 簇)
The importance of packing of DNA into chromosomes • Chromosome is a compact form of the DNA that readily fits inside the cell • To protect DNA from damage • DNA in a chromosome can be transmitted efficiently to both daughter cells during cell division • Chromosome confers an overall organization to each molecule of DNA, which facilitates gene expression as well as recombination
Proteins in chromosome (1) • Half of the molecular mass of eukaryotic chromosome is protein • In eukaryotic cells a given region of DNA with its associated proteins is called chromatin • The majority of the associated proteins are small, basic proteins called histones. • Other proteins associated with the chromosome are referred to as non-histone proteins, including numerous DNA binding proteins that regulate the transcription, replication, repair and recombination of DNA.
Proteins in chromosome (2) • Nucleosomes: regular association of DNA with histones to form a structure effectively compacting DNA
CHAPTER 7: Chromosomes, chromatin, and the nucleosome • What is the cost (challenge) of compaction of DNA into chromosome? • [De-compaction is required when DNA needs to be accessible by other proteins for cellular activities such as replication, transcription. ] • 2. How the challenge could be resolved? • [The interaction between DNA and histones is dynamic, chromosome remodeling, chromosome modification] • 3. What are the advantage of the challenge? • [Add a large layer of gene regulation]
CHAPTER 7: Chromosomes, chromatin, and the nucleosome OUTLINE • Chromosome sequence & diversity • Chromosome duplication & segregation • The nucleosome • Higher-order chromatin structure • Regulation of chromatin structure • Nucleosome assembly
CHAPTER 7: Chromosomes, chromatin, and the nucleosome Chromosome sequence & diversity Chromosomes • Shape: circular or linear • Number in an organism is characteristic • Copy: haploid, diploid, polyploid Genomes
Chromosome sequence & diversity Difference in the structures of eukaryotic and prokaryotic cells is akeyto better understand the molecular processes of genome maintenance and expression, as well as the differences in these processes between eukaryotes and prokaryotes
Figure 7-1* Chromosome sequence & diversity
Chromosome sequence & diversity Genome & the complexity of the organism • Genome size: the length of DNA associated with one haploid complement of chromosomes • Gene number: the number of genes included in a genome • Gene density: the average number of genes per Mb of genomic DNA See Table 7-2 to find the relationship
Chromosome sequence & diversity Genes make up only a small proportion of the eukaryotic genome • Increases in gene size: (1) increase in the sequence of regulatory sequence; (2) presence of introns (splicing) • Increases in the DNA between genes (intergenic sequences): (1) unique; (2) repeated See Figure 7-2, 3, 4, 5; Table 7-3
CHAPTER 7: Chromosomes, chromatin, and the nucleosome Chromosome duplication & segregation (1) Critical DNA elements • Origins of replication • Centromeres • Telomeres These elements are not involved in gene expression
Chromosome duplication & segregation Origins of replication Sites at which the DNA replication machinery assembles to initiate replication; required fro replication • 30-40 kb apart on each eukaryotic chromosome • Only one origin for prokaryotic chromosome
Chromosome duplication & segregation Centromeres Required for the correct segregation of the chromosomes after replication • Direct the formation of kinetochore (an elaborate protein complex) essential for chrom. segregation • One chromosome, one centromere • The size varies (200 bp- >40 kb) • Composed of largely repetitive DNA sequences
Figure 7-6 Centromeres, origin of replication and telomere are required for eukaryotic chrom. maintenance
(2) Eukaryotic chromosome duplication & segregation occur in separate phases of the cell cycle Chromosome duplication & segregation Cell cycle: a single round of cell division Mitotic cell division: the chrom. Number is maintained during cell division
Figure 7-11 The events of M phase Mitotic spindle
Chromosome duplication & segregation (3) Chromosome structure changes as eukaryotic cells divide M phase: condensed state, completely disentangled from each other G1, S, G2 phases: diffused, significantly less compact. The structure of chrom. changes, e.g. DNA replication requires the nearly complete disassembly and reassembly of the proteins associated with each chromosome
Figure 7-13 Changes in chromatin structure Chromosome condensation REMEMBER: chromosome is a consistently changing structure (dynamics)
Chromosome duplication & segregation (4) The gap phase of the cell cycle allow time to prepare for the next cell cycle stage while also checking that the previous stage is finished correctly. Think of the regulatory mechanisms might be involved to CHECK the cell condition
Chromosome duplication & segregation (5) Different levels of chromosome structure can be observed by microscopy
Figure 7-13 Forms of chromotin structure seen in EM (electron microscopy)
CHAPTER 7: Chromosomes, chromatin, and the nucleosome The nucleosome Nucleosome & histone structures
(1) Nucleosomes are the building blocks of chromosomes The nucleosome • The nucleosome is composed of a core of eight histone proteins and the DNA (core DNA, 147 bp) wrapped around them. The DNA between each nucleosome is called a linker DNA. Each eukaryote has a characteristic average linker DNA length (20-60 bp)
Figure 7-18 DNA packaged into nucleosome Six-fold DNA compaction
(2) Histones are small, positively charged (basic) proteins • Five abundant histones are H1 (linker histone, 20 kd), H2A, H2B, H3 and H4 (core histones, 11-15 kd). • The core histones share a common structural fold, called histone-fold domain • The core histones each have an N-terminal “tail”, the sites of extensive modifications The nucleosome
Figure 7-19 The core histones share a common structural fold (1) (2)
(3) Many DNA sequence-independentcontacts (?) mediate interaction between the core histones and DNA The nucleosome Figure 7-25
(4) The histone N-terminal tails stabilize DNA wrapping around the octamer Figure 7-26 The histone tails emerge from the core of the nucleosome at specific positions, serving as the grooves of a screw to direct the DNA wrapping around the histone core in a left-handed manner.
CHAPTER 7: Chromosomes, chromatin, and the nucleosome Higher-order chromatin structure How does it form?
(1) Histone H1 binds to the linker DNA between nucleosome, inducing tighter DNA wrapping around the nucleosome Higher-order chromatin structure Figures 7-28, 29
(2) Nuclear arrays can form more complex structures: the 30-nm fiber (“zigzag model”) Higher-order chromatin structure Figures 7-30 (40-fold compaction)
(3) Further compaction of DNA involves large loops of nucleosomal DNA Higher-order chromatin structure • Additional 103-104-fold compaction is required, but the mechanism is unclear • The nuclear scaffold model is proposed
Figures 7-32 The higher- order structure of chromatin. (a) A transmission electron micrograph, (b) A model
Higher-order chromatin structure (3) Histone variants alter nucleosome function • Several histone variants are found in enkaryotes • This variants can replace one of the 4 standard histones to form alternate nucleosomes
Figures 7-33 Alteration of chromatin by incorporation of histone variants CENP-A is associated with the nucleosomes containing centromeric DNA
CHAPTER 7: Chromosomes, chromatin, and the nucleosome Regulation of chromatin structure How?
Regulation of chromatin structure The interaction of DNA with the histone octamer is dynamic • There are factors acting on the nucleosome to increase or decrease the dynamic nature • The dynamic nature of DNA-binding to the histone core is important for access of DNA by other proteins essential genome expression etc.
Regulation of chromatin structure Nucleosome remodeling complexes facilitate nucleosome movement • A large protein complexes facilitate changes in nucleosome location or interaction with the DNA using the energy of ATP hydrolysis.