Comprehensive Molecular Biology and DNA Synthesis Guide
550 likes | 686 Vues
Explore the fundamentals of molecular biology, DNA structure, replication, and repair processes. Learn about nucleotide components, polymerases, chromatin organization, and DNA replication mechanisms in this detailed educational resource.

Comprehensive Molecular Biology and DNA Synthesis Guide
E N D
Presentation Transcript
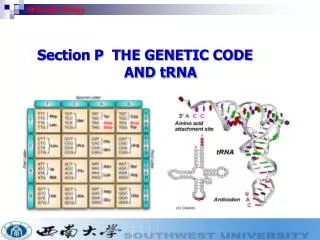
Molecular Biology The DNA
Intended learning outcomes • Describe the base, sugar and phosphate moieties of nucleotides. • Describe phosphodiester bond and the concept of chain polarity and the 3' and 5' ends of polynucleotides. • Describe the main features of the double helical Watson Crick model of DNA. • Explain the implications of the antiparrallel complementary nature of DNA. • Describe 4 types of template directed nucleic acid polymerases. • Describe the structure of chromatin and chromosomes. • Describe the general features of DNA replication. • Describe the role of DNA polymerases in replication. • Explain the attainment of fidelity of DNA replication. • Describe the function of proteins and enzymes at the replication fork. • Outline damage to DNA and its repair
DNA synthesis Where in the cell cycle does DNA synthesis occur
From nucleotides to replication and maintenance of the genome • The structure and nomenclature of mono- and poly-nucleotides • Understanding DNA structure • The packaging of DNA into chromosomes • Nucleic acid polymerases and the mechanism of DNA replication • DNA damage and repair
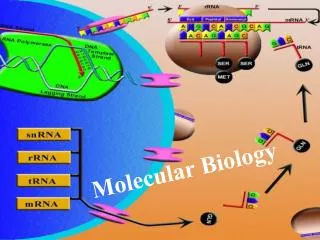
THE CENTRAL DOGMA Transcription Translation RNA Protein DNA DNA replication
A nucleotide consists of a sugar, base and phosphate group 5’ O P BASE 1’ 4’ SUGAR 3’ 2’
The difference between a nucleotide and nucleoside is the presence of phosphate groups
Some examples • If base is adenine and sugar deoxyribose and there are 3 phosphates – DEOXY ADENOSINE TRIPHOSPHATE, dATP • If base is adenine and sugar ribose and there is 1 phosphate – ADENOSINE MONOPHOSPHATE, AMP
Polynucleotides are nucleotide chains joined by phospho-diester bonds 3’ to 5’ phospho- diester bond
5’ 3’ Major groove Minor Groove 3’ 5’ Watson-Crick model of DNA structure
The purines and pyrimidines can interact with one another to form base pairs
DNA is a helical structure of two strands of polynucleotides 5’ 3’ 3’ 5’
The principal features of the structure of DNA • DNA consists of two helical polydeoxyribonucleotide strands • The sugar-phosphate backbones of the strands are antiparallel; they run 3’―5’ and 5’―3’ • The strands are joined non-covalently by hydrogen bonding between bases • Bases always pair according to the rule A=T and G≡C • DNA normally occurs with the following dimensions: 10bp per turn of the helix 3.4nm per turn of the helix 2.0nm diameter • Each nucleotide carries a negative charge on the phosphate
What differentiates species is not the way DNA is packaged……. Fruit fly: 1.7x108bp; 4 chromosomes Yeast: 1.4x107bp; 16 chromosomes Human: 3x109bp; 23 chromosomes A 2% difference in genome sequence results in different species
DNA Replication rates in E.Coli (Bacteria) • In E.coli cells, it takes 42 minutes to replicate a circular chromosome that has 4,639,221bp and is about 4.1mm in length • Since the chromosome is duplicated from one origin by two growing forks, the rate of fork movement is about 1000bp per second per fork.
DNA replication rates in humans • The rate of fork movement in human cells is 100bp per second per fork. • The entire human genome of 3,000,000,000 bp replicates in about 8 hours, suggesting around 1000 forks • However there are about 10,000-100,000 replicons, each of which operates for only part of the 8 hour period
DNA Replication • Replication of bacterial genomes occur at a single origin of replication • There are unique sequences at the origin of replication • Several proteins recognize the origin and separate the DNA strands
A replication fork of E.Coli Direction of fork movement
DNA helicase unwinds the DNA ahead of the DNA replication fork • Single stranded DNA binding proteins (SSB) are also at the replication forks to protect the DNA from cleavage • A special type of RNA polymerase, primase, makes short 10 to 20 nucleotide chains of RNA that are complementary to one of the template DNA strands • Nascent DNA is covalently linked to the short stretch of RNA • Therefore RNA primes the synthesis of DNA
DNA polymerases synthesise DNA • DNA polymerases carry out the following reaction: (dNMP)n+dNTP = (dNMP)n+1 + PPi • Synthesise chains ONLY in a 5’ to 3’ direction • DNA polymerases synthesise DNA with a high degree of fidelity • DNA is read in a 5’ to 3’ direction • There are three DNA polymerase I,II and III. They can synthesize DNA and proof read using a 3’ to 5’ exonuclease activity
DNA polymerases I, II and III are proof reading 3’ to 5’ exonucleases
Only DNA polymerase I is an error correcting 5’ to 3’ exonuclease. This activity is essential for removing the RNA from the DNA
Summary of replication in E.Coli • Origin of replication is recognized by a number of proteins • These include DNA helicase and single stranded DNA binding proteins (SSB) • The result of this recognition is the unwinding of the double stranded DNA helix • Primase synthesises a 10-20 base RNA strand in the 5’ to 3’ direction that is complementary to the DNA • This is then extended by DNA polymerase III
At the replication fork there is a leading and a lagging strand. Leading strand presents no problems as it can be synthesised in the 5’ to 3’ direction • On the lagging strand the DNA is synthesised as Okazaki fragments which are around 1,000 to 2,000 bases in length • DNA polymerase I removes the RNA primer and fills in the gap between Okazaki fragments • The Okazaki fragments are then joined together by DNA ligase • Any mistakes in incorporation of nucleotides can be repaired by a 3’ to 5’ exonuclease activity
Condensation of DNA into chromosomes • The length of DNA presents a problem • DNA in eukaryotic chromosomes is tightly bound to basic proteins called histones • Histones consist of half of the mass of chromosomes, the other half being DNA • The nucleoprotein chromosomal content is termed chromatin • There are five types of histones: H1, H2A, H2B, H3, H4 • Histones are extremely basic; about 1 in 4 amino acids are basic (arginine or lysine) • This basic content gives histones an overall positive charge allowing for interaction with negatively charged DNA
A nucleosome core consists of 140bp of DNA wound around a histone octamer
Chromatin is made up of repeating units of 200bp; 140bp wound round the histone core (the nucleosome) with additional linker DNA • It is postulated that the nucleosomes themselves form a helical array making a solenoidal structure • Solenoids attach to a nuclear matrix (a protein scaffold) • This scaffold is then itself condensed, again potentially into a helical structure • All of this condensation allows the DNA to pack into chromosomes that we find in cells
Formation of nucleosomes into solenoids Nucleosome
Not all DNA is equally condensed in the nucleus • Euchromatin; dispersed appearance and occupies most nuclear volume • Heterochromatin; densely packed
Sources of DNA damage • Spontaneous damage loss of purine base loss of pyrimidine base deamination • Damage induced by external agents e.g. chemicals, U.V. light (sunshine!), tobacco
Spontaneous Damage • Loss of purines and pyrimidines: (hydrolytic loss of the base from the sugar) • Mis-incorporation of a nucleotide during replication; e.g. a T is incorporated next to a G. • Deamination (hydrolytic loss of the NH2 group from a base)– particularly with cytosine changing it to uracil. This is potentially mutagenic as uracil will behave like thymine upon DNA replication
External Damage • Physical agents including ultraviolet (UV) light (causes pyrimidine dimer formation) and X-rays (backbone breakage) resulting in inhibition of DNA replication and loss of genome integrity • Chemical agents such as nitrous acid (deamination) resulting in mutation, and N-alkylation resulting in depurination
Consequences of DNA Damage • Damage causes change in non-essential DNA • The damage causes a change in an essential part of the genome but does not alter the cell’s information • The damage is repaired before it can exert any harmful effect • Mutations in essential coding regions of the genome that are inherited by the daughter cells • Results in a change in protein sequence and therefore potentially function - mutation • Can result in several diseases, including cancer
Mutated DNA can be repaired • Most DNA repair involves the following steps • Recognition; this is detection of the damage • Incision; cutting into the damaged DNA • Excision; cutting out the damaged part • Polymerisation; gap filled by DNA polymerase • Religation; joining of the resulting nick carried out by DNA ligase
G - C Replication Deamination G - U G - C A - U Example: how deamination results in mutation