Understanding RNA Transcription and Its Modifications in Protein Synthesis
This detailed exploration of protein synthesis focuses on RNA transcription and its critical role in the process. It covers the initiation, elongation, and termination of transcription, highlighting the functions of RNA polymerase and the significance of promoter regions. Additionally, it delves into post-transcriptional modifications of precursor mRNA in eukaryotes, including 5’ cap addition, poly-A tail addition, and intron splicing. These modifications ensure the stability and functionality of mature mRNA, allowing it to be properly translated into proteins.


Understanding RNA Transcription and Its Modifications in Protein Synthesis
E N D
Presentation Transcript
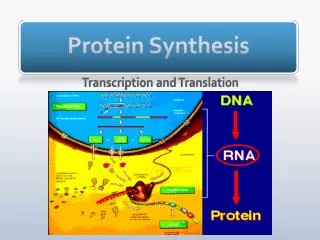
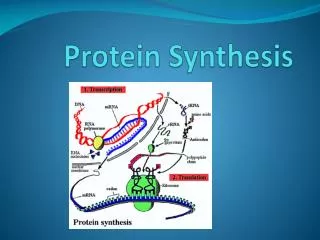
PROTEIN SYNTHESIS THE DETAILS
Making RNA from DNA Transcription
RNA Pg 251
Comparing DNA & RNA Pg 251
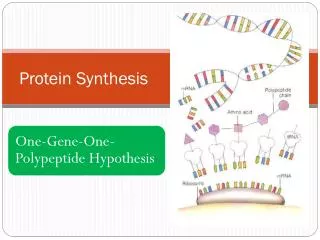
RNA • Different types of RNA • All are produced from a DNA template in the nucleus • Classified according to their function in the cell
DNA Template Strand Transcription mRNA Translation Protein A T G G A C T T A 5’ 3’ 5’ U A C C U G A A U 3’ Amino acid Amino acid Amino acid
TRANSCRIPTION: the details INITIATION • RNA polymerase binds to promoter (sequence of DNA upstream of the gene to be transcribed) – opens up the double helix • promoters contain base pair patterns rich in A’s and T’s ….Why???
DNA strand is unwound, double helix disrupted…exposing template strand
ELONGATION • RNA polymerase starts building mRNA in the 5’ to 3’ direction • process similar to DNA replication – except… • no primer is used • only 1 strand of DNA is used -the template strand or the antisense strand
DNA template is used to synthesize mRNA in 5’ to 3’ (uracil complements adenine)
DNA that has already been transcribed rewinds into double helical form
ELONGATION • unused DNA strand is called the coding strand or the sense strand ***mRNA is complementary to the template strand and identical to the coding strand (except it contains U)
TERMINATION • Specific nucleotide sequences in the template serve as a signal to stop transcription • mRNA dissociates from the DNA template • RNA polymerase is free to transcribe another gene.
mRNA MODIFICATIONS in Eukaryotes • Convert precursor mRNA (pre-mRNA) to mature mRNA 1) 5’ cap added • 7-methyl guanosine added to 5’ end • Cap is recognized by the protein synthesis machinery
mRNA MODIFICATIONS in Eukaryotes 2) Poly-A tail added • The addition of a series of A nucleotides to the 3’ end of the pre-mrNA • Makes mRNA more stable, allows it to exist longer in cytoplasm animation
mRNA MODIFICATIONS in Eukaryotes 3) Introns cut out • eukaryotic genes have coding regions called exons, and non-coding regions called introns • Splicing – intron sequences are removed and the exons are joined together to form the mature mRNA
mRNA MODIFICATIONS in Eukaryotes 3) Introns cut out • snRNP’s (particles of snRNA and protein) recognize regions where exons and introns meet, and bind to those areas • snRNP’s interact with other pr- forming a larger spliceosome complex that removes introns
mRNA MODIFICATIONS in Eukaryotes 3) Introns cut out • for the expression of most genes, all of the exons are spliced together • BUT, sometimes only certain exons are used to form a mature RNA transcript (alternative splicing)
mRNA MODIFICATIONS in Eukaryotes 3) Introns cut out • Alternative splicing allows for one gene to code for more than one protein • Certain cell types are able to produce forms of a pr- that are specific for that cell
Unlike DNA replication, no quality control in mature RNA, therefore, there are more errors made during transcription than DNA replication….BUT…
Since a single gene is transcribed many times to produce 100’s of transcripts, errors are not as detrimental. • Errors in mRNA result in a protein being made that is susceptible to degradation.