Apoptosis
PRESENTED BY DOCTOR DEATH. Apoptosis. Learning Objectives To Die For. To understand the proteins involved in apoptosis. Know the regulation of apoptotic signaling pathways. Importance of apoptosis in the immune system and diseases. Apoptosis (Programmed cell death).

Apoptosis
E N D
Presentation Transcript
PRESENTED BY DOCTOR DEATH Apoptosis
Learning Objectives To Die For • To understand the proteins involved in apoptosis. • Know the regulation of apoptotic signaling pathways. • Importance of apoptosis in the immune system and diseases.
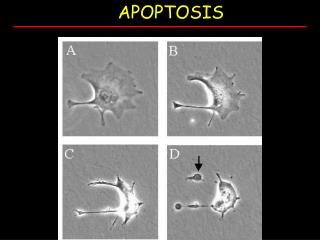
Apoptosis (Programmed cell death) • Active process of cell collapse • Requires ATP • Morphological Features • Membrane blebbing • Chromatin condensation • Protein fragmentation • Apoptotic bodies (engulfed by phagocytosis)
Apoptosis • Apoptosis is conserved from worms to mammals • In humans 10 billion cells undergo apoptosis to maintain number of new cells generated each day • In the immune system development, over-production of cells arise followed by death of these cells that fail to have productive antigen specifications • Apoptosis is necessary to purge the body of pathogens invaded cells and eliminate activated or autoimmune cells.
Apoptosis • Apoptosis needs to be tightly regulated since too little or too much cell death may lead to developmental defects, autoimmune disease, or cancer such as leukemia. • There are now over 50,000 paper on apoptosis • Many companies are developing therapeutic drugs targeted against apoptosis for treatment of diseases.
Morphological changes during apoptosis Department of Biochemistry and Immunology at St.George's Hospital Medical School, London.
C. Elegans • First organism in which regulation of apoptosis was discovered. • C. elegans is special since every lineage of cells is known and ultimate fate of each cell is known. • Genetic analysis by elimination of genes revealed four genes essential for apoptoiss • CED 3 • CED 4 • CED 9 • Egl 1
CED 3 CED 4 CED 9 Egl 1 APOPTOSIS
CED 3 • CED 3 is a cystiene protease homologous to proteases in mammals called caspases. • Caspases cleave specific proteins • Proteolysis is irreversible • Regulation involves - substrate availability • - catalytic activity • - protein turnover
Activation involves proteolytic cleavage of the subunits into a tetramer complex Active caspase Caspases Expressed as a proenzyme Prodomain (30-50kDa) Large subunit (20kDa) Small subunit (10kDa)
Caspases • Activated caspases cleave at four amino acids with aspartic acid required at the amino terminal end. • Consensus sites for caspases • Caspase 8 (L/V/D)EXD • Caspase 9 (I/V/L)EHD • Caspase 3 DEXD • Caspase 1 (W/Y/F)EHD
Classifications of Caspases Caspase 8 Caspase 10 Initiation Caspases Caspase 9 Caspase 2 Caspase 1 Intermediate Caspases Caspase 4 Capsase 5 Caspase 3 Terminal (Effector) Caspases Caspase 6 Caspase 7
Caspase • Substrates: • Caspases cleavage many proteins that can be categorized into four general groups • Nuclear proteins • Structural proteins • Signaling proteins • Caspases
Department of Biochemistry and Immunology at St.George's Hospital Medical School, London.
Caspases • Regulation of Effector Caspases • Cascade model • Initiator caspases activate effector caspases through cleavage. • Regulation of Initiator Caspases • Activation involves co-factors • Binding of co-factors triggered by a pro-apoptotic signal • Mediated through two distant structural motifs • Prodomain of caspases and its corresponding co-factor • Caspase 8 pro-domian binds to the adaptor protein FADD through a death effector domain (DED) • Caspase 9 binds to a co-factor through the caspase recruitment domain (CARD) found in Apaf1
Caspases • How do co-factors activated caspases? • Induced proximity or oligomerization model • Based on • Low but detectable levels of activity • Dimerization required for activation • Over-expression and cross-linking leading to activation • Caspases are latant and co-factors bring two or more caspases together allowing for intermolecular auto-proteolytic activation • Faciliated autocataylsis model • Caspase precursors are present in complexes • Co-factors facilitate activation by conformational changes either directly or by removing an inhibitory protein. • Compartmentaliztion • A co-factor is sequester in a different location in the cells than the caspase • Inhibitors binding to caspases prevent binding of co-factors.
CED 4 CED 3 APOPTOSIS
Co-Factor CED 4 • CED 4 contains a CARD domain and is homologous to Apaf1. • Afap1 is localized to the cytoplasmic side of the mitochondria. • Binds to caspase 9 through its CARD domain • Requires cytochrome c and ATP to bind to caspase 9 and activate its activity. • Caspase 9 cleavage might not be necessary for its activation.
Caspase 2 has co-factors! Rastogi et al EXCLI Journal 2009
Co-Factors CED4 • Besides APAF-1, another co-factor exists called FAS associated death domain (FADD). • Binds caspase 8 and caspase 10 • FADD inducibly associates with activated death receptors causes caspase trans-cleavage and activation
Types of Apoptotic Signaling Pathways • Extrinsic Apoptotic pathway • Death receptor activation • Intrinsic Apoptotic pathway • Mitochondria regulated
Death Receptors • Tumor necrosis factor receptor (TNFR) family is involved in the induction of apoptosis. • Members contain a five cysteine rich repeats in the extracellular domain and a death domain in the cytosolic tail. • Ligation of these receptors causes rapid induction of apoptosis.
? TRADD FADD FADD TRADD FADD FADD FADD FADD Caspase8 Caspase 8 Caspase 8 Caspase8 Caspase 8 Caspase 8 Active caspase 8 APOPTOSIS Death Receptor Family FASL TRAIL TNF FAS DR4/5 TNFR
Department of Biochemistry and Immunology at St.George's Hospital Medical School, London.
TRAIL DR4/ DR5 TRAIL DR4/ DR5 DR4/ DR5 DD DD FADD DD pro-caspase 8 Caspase 8 tBid • R.O.S. • DYM • Release of Smac/Diablo Signal Transduction Upstream of Mitochondria TRAIL Cell Membrane Cytosol
FAS Receptor • Expression is increased following cytokine, or lymphocyte activation. • Ligand for FAS receptor is FAS ligand that is a type II transmembrane molecule and its expression is tightly controlled. • FAS is trimerized following FAS ligand binding and is required for its activation. • A complex of proteins associate with the receptor called the death inducing signaling complex (DISC).
FAS Receptor • In Type I • Caspase 8 cleaves and activates caspase 3 • In Type II • Caspase 8 cleaves BID and releases cytochrome c from the mitochondria. • DISC formation is limited and BID cleavage serves to amplify the apoptotic signal. • FASL membrane bound vs soluble might determine type of FAS activation?
Mitochondria Bid tBid caspase-3 APOPTOSIS Beyond Caspase 8 Activation FASL FAS TYPE II FADD Procaspase-8 Initiator Caspase Active caspase-8 TYPE I Effector Caspase
TRAIL (a death receptor ligand) • Binds to death receptor 4 and 5. • Activates caspase 8 through binding of FADD to receptors. • Synergy between TRAIL and chemotherapeutic drugs in cancer cells.
Control EGF TRAIL Pro-Caspase 8 BID MKP-1
Decoy Receptors: STOP THE DESTRUCTION • Expressed on the cell surface and bind to death receptor ligands such as TRAIL • Fail to recruit FADD to the receptor and thereby fails to activate caspase 8. • Blocks death receptor induced apoptosis by sequestering its ligand.
FASL TRAIL DcR1/2 DcR3 FLIP FLIP Active caspase 8 FASL TRAIL FAS DR4/5 ? FADD FADD FADD FADD Caspase8 Caspase 8 Caspase 8 Caspase8 APOPTOSIS
Intrinsic Apoptotic Pathway • Requires mitochondria • Three main events happen in the mitochondria to induce apoptosis • Loss of Dym • Increase reactive oxygen species (ROS) • Release of mitochondrial proteins
Release of Mitochondrial Protein • During apoptosis, cytochrome c is released from the mitochondria. • It binds to caspase 9/Apaf1 complex thereby activating caspase 9 activity • This complex is called the apoptosome (same concept as the DISC). • Leads to effector caspase activation and apoptosis.
Release of Mitochondrial Protein • Smac/Diablo is another protein release from the mitochondria during apoptosis. • Binds to inhibitor of apoptosis proteins (IAP) • IAP constitutively bind to caspases (both initiator and effector). • Smac/Diablo sequestors IAPs away from caspases allowing for caspase activation • Takes the brakes off the apoptotic pathway.
CELL Apoptotic Signal IAPs Healthy Death
Smac/Diablo Omi/HtrA2 FADD APAF1 Caspases Caspases Caspases Caspases Caspases Caspases Caspases GAS CELL CELL CELL CELL CELL CELL CELL Caspases CELL Apoptotic Signal Healthy Death
Other Proteins Released • Apoptosis inducing Factor (AIF) is released from the mitochondria. • Translocates to the nucleus and participates in DNA fragmentation. • Nuclease G is a Dnase that also translocates to the nucleus and degrades DNA.
The Mitochondria and Apoptosis • Different pro and anti-apoptotic signals converge at the mitochondria. • Apoptotic signals activate pro-apoptotic Bcl-2 family members such as Bid, Bax, and Bak. Survival signals work through the action of anti-apoptotic Bcl-2 family members such as Bcl-2, and Bcl-XL. • Key apoptotic events include a loss in the mitochondrial membrane potential DyM, the production of reactive oxygen species R.O.S., and the release of mitochodrial proteins such as Smac/Diablo and cytochrome c.
Role of Mitochondria • Two well defined compartments in the mitochondria • Inner membrane where the electron transport chain is found • Outer membrane that controls solutes and ions into the mitochondria. • Permeable transition pore (PT pore) controls the movement of ions and solutes in and out of the mitochondria. • Changes in this pore during apoptosis reduces the electrochemial gradient and increases reactive oxygen species (ROS). • PT pore might also control release of proteins?
PT Pore Cyclosporin A Hexokinase PBR VDAC CK ANT CycD ? DYm Cytochrome c Smac/Diablo AIF Omi/Hrti Nuclease G - - - - - +++++ Inner Membrane Outer Membrane Mitochondria
CED 9 Egl 1 CED 3 CED 4 APOPTOSIS