3.5 (SL)/ 7.3 (HL): Transcription
620 likes | 930 Vues
3.5 (SL)/ 7.3 (HL): Transcription. Compare the structure of DNA and RNA. Compare the processes of transcription and translation. DNA transcription (higher level). Let’s begin with some animations…. from the Wellcome trust From PBS.

3.5 (SL)/ 7.3 (HL): Transcription
E N D
Presentation Transcript
Let’s begin with some animations… • from the Wellcome trust • From PBS
Exceptions to the central dogma of genetics 1: retroviruses (thanks Max)
Prion protein replication Prions are proteins that propagate themselves by making conformational changes in other molecules of the same type of protein
And finally….thanks, Ilona! • exceptions
7.3.1: State that transcription is carried out in a 5’ – 3’ direction
Remember that BOTH transcription AND translation are carried out in a 5’ to 3’ direction
7.3.2: Distinguish between the sense and antisense strands of DNA • Sense strand is the genetic code • Sense strand is not copied • Antisense strand is the template strand • Antisense strand is used for transcription • Antisense strand is complementary to sense strand
Sense and antisense DNA • SENSE strand has same base sequence as transcribed mRNA, except that T is replaced by U • RNA polymerase enzyme adds RNA nucleotides to sugar-phosphate background in 5’ – 3’ direction
Where does transcription begin? • Promotor region: contains ‘promotor’ sequence: specific DNA recognised by activator proteins wh
Promotor Region • Upstream of transcription site • Contains specific DNA (TATAAA boxes) recognised by activation factors which recruit RNA polymerase • 100 – 100 bases long • Very diverse in eukaryotic DNA
Coding Region • Provides specific triplet code for specific protein • Coding region produces mRNA which will be modified after transcription • mRNA produced by RNA polymerase
Terminator Region • Site signals the end of DNA transcription • Triplet code makes both RNA polymerase AND mRNA strand fall off the template • Fold up mRNA strand to unlock both mRNA and polymerase
Process of transcription • Animation of transcription • more complete animation of transcription
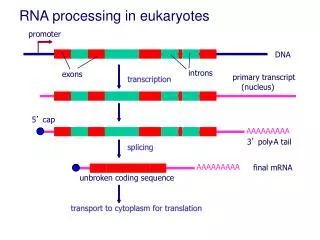
Post-translational modification • Eukaryotes only • mRNA is processed in the nucleus before exportation to the cytoplasm • Introns are removed (spliced) • Mature mRNA is exported to the cytoplasm • EXONS are joined together
Post-translational modification - splicing • Animation on RNA splicing
Translation! Let’s start with some animations… From John Kyrk From Wiley From Harvard
Ribosomal RNA Ribosomal structure form the PDB
Transfer (tRNA) • ‘Adaptor molecules’: one end can read the triplet code on mRNA (anticodon), one end(3’) attaches to a specific amino acid (Crick, 1958) • Each tRNA has a specific shape, determined by looping of helical sections • Each tRNAspecific shape fits one of 20 aa-tRNA synthase enzymes
Structure of tRNA • Amino acid attachment site (3’) (acceptor loop/stem) :always CCA • Complementary base pairing (H-bonds) • Non-base pairing loops (7 or 8 bases) • Anticodon (3 bases): atttaches to mRNA codon • Mammals have 150 tRNA molecules
Activation of tRNA • Activation of tRNA
How does the correct amino acid link to tRNA? • The shape of each tRNA is different, defined by folding of the loop and helical structures RNA folding • The shape of the tRA determines which of 20 specific amino-acyl tRNA synthase enzymes it attracts • ATP is needed to attach the amino acid to the 3’ CCA end of the tRNA • Each amino acid has >1 tRNA molecule (degenerate code)
Structure of a ribosome (ribozymes) • Ribosomes are actually ribozymes • They catalyse translation of mRNA into a polypeptide • The substrate is mRNA • Each ribosome is multifunctional – it is not used up, and catalyse translation of many different mRNA codes













![[V]. Process of Transcription and Transcriptional Control of Gene Expression](https://cdn2.slideserve.com/5058527/v-process-of-transcription-and-transcriptional-control-of-gene-expression-dt.jpg)