### Advanced Hidden Markov Models for Sequence Alignment: Theory and Applications ###
690 likes | 842 Vues
This text elaborates on the application of Hidden Markov Models (HMMs) for multiple sequence alignment without the need for indel penalties. It emphasizes the utility of HMMs to synthesize experimentally derived information and adjust parameters that reflect observed variations across at least 20 sequences. The document explores evolutionary aspects of sequences, including the introduction of mutations and divergence over time, and highlights how HMMs can act as intermediates to uncover relationships among sequences, enhancing our understanding of sequence evolution and protein structure. ###

### Advanced Hidden Markov Models for Sequence Alignment: Theory and Applications ###
E N D
Presentation Transcript
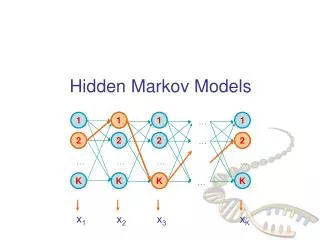
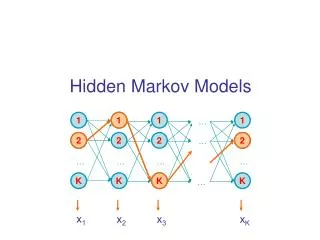
Hidden Markov Models • Probabilistic model of a Multiple sequence alignment. • No indel penalties are needed • Experimentally derived information can be incorporated • Parameters are adjusted to represent observed variation. • Requires at least 20 sequences
The Evolution of a Sequence • Over long periods of time a sequence will acquire random mutations. • These mutations may result in a new amino acid at a given position, the deletion of an amino acid, or the introduction of a new one. • Over VERY long periods of time two sequences may diverge so much that their relationship can not see seen through the direct comparison of their sequences.
Hidden Markov Models • Pair-wise methods rely on direct comparisons between two sequences. • In order to over come the differences in the sequences, a third sequence is introduced, which serves as an intermediate. • A high hit between the first and third sequences as well as a high hit between the second and third sequence, implies a relationship between the first and second sequences.Transitive relationship
Introducing the HMM • The intermediate sequence is kind of like a missing link. • The intermediate sequence does not have to be a real sequence. • The intermediate sequence becomes the HMM.
Introducing the HMM • The HMM is a mix of all the sequences that went into its making. • The score of a sequence against the HMM shows how well the HMM serves as an intermediate of the sequence. • How likely it is to be related to all the other sequences, which the HMM represents.
B M1 M2 M3 M4 E Match State with no Indels MSGL MTNL Arrow indicates transition probability. In this case 1 for each step
B M1 M2 M3 M4 E Match State with no Indels MSGL MTNL S=0.5 T=0.5 M=1 Also have probability of Residue at each positon
B M1 M2 M3 M4 E Typically want to incorporate small probability for all other amino acids. MSGL MTNL S=0.5 T=0.5 M=1
B M1 M2 M3 M4 E Permit insertion states MS.GL MT.NL MSANI I0 I1 I2 I3 I4 Transition probabilities may not be 1
B M1 M2 M3 M4 E Permit insertion states MS..GL MT..NL MSA.NI MTARNL I0 I1 I2 I3 I4
DELETE PERMITS INCORPORATION OF LAST TWO SITES OF SEQ1 MS..GL-- MT..NLAG MSA.NIAG MTARNLAG AA GN IL ST A M D1 D2 D3 D4 D5 D6 D7 I7 I0 I1 I2 I3 I4 I5 I6 M4 E M2 M3 M5 M6 B M1 M7 G
D1 D2 D3 D4 D5 D6 I0 I1 I2 I3 I4 I5 I6 B M1 M2 M3 M4 M5 M6 E • The bottom line of states are the main states (M) • These model the columns of the alignment • The second row of diamond shaped states are called the insert states (I) • These are used to model the highly variable regions in the alignment. • The top row or circles are delete states (D) • These are silent or null states because they do not match any residues, they simply allow the skipping over of main states.
Dirichlet Mixtures • Additional information to expand potential amino acids in individual sites. • Observed frequency of amino acids seen in certain chemical environments • aromatic • acidic • basic • neutral • polar
STRUCTURES a helix b sheet coils turns Structures are used to build domains.-Legos of evolution
Ramachandran plot for Glycine Areas not permitted for other amino acids Psi Angles Phi angles
Introduction to Protein Structure, Branden and Tooze Garland Publishing Co.1991 p.13
From: http://bioweb.ncsa.uiuc.edu/~bioph254/Class-slides/Lect12/figure13.html
Longitudinal and Transverse image of alpha helix From: http://bioweb.ncsa.uiuc.edu/~bioph254/Class-slides/Lect12/figure14.html
Turn connecting two helices Introduction to Protein Structure, Branden and Tooze Garland Publishing Co.1991 p. 17
Proline • Because of its structure, proline is typically excluded from a helices except in the first three positions at the amino end.
b Structure b strand - single run of amino acids in b conformation b sheet- multiple b strands which are hydrogen bonded to yield a sheet like structure. b bulge - disruption of normal hydrogen bonding in a b sheet by amino acid(s) that will not fit into the sheet -for example: proline
b sheets- Parallel Introduction to Protein Structure, Branden and Tooze Garland Publishing Co.1991 p.17.
b sheet - longitudinal and transverse view. Side chains stick “out” http://bioweb.ncsa.uiuc.edu/~bioph254/Class-slides/Lect12/figure22.html
Six classes of structure • Class a- bundled a helices connected by loops. • Class b- sandwich or barrel comprised entirely of b sheets typically anti-parallel. • Class a / b mainly parallel b sheets with intervening a helices. • Class a + b - segregated a helices and anti-parallel b sheets • Multi-domain • Membrane proteins
Endonuclease Class a + b
Rhodopsin 7TM proten
Common Hairpin Loop between two b Strands Introduction to Protein Structure, Branden and Tooze Garland Publishing Co.1991 p. 17
Turn - short, regular loop. • Difference in frequency of amino acids at positions 1-4 of the turn. • Coils (not coiled coil) • Random turns or irregular structure.
Disulfide bridges • Crosslink of two cysteine residues. • Distance between sulfur = 3 Angstroms.
Coiled coil -two a helices bundled side by side From: http://catt.poly.edu/~jps/coilcoil.html
a,d are internal, remaining amino acids are solvent exposed From: http://catt.poly.edu/~jps/coilcoil.html
Coiled Coil • Two or more adjacent a helices
Potential Residues involved in Coiled Coil MMFPQSRHSGSSHLPQQLKFTTSDSCDRIKDEFQLLQAQYHSL KLECDKLASEKSEMQRHYVMYYEMSYGLNIEMHKQAEIVKR LNGICAQVLPYLSQEHQQQVLGAIERAKQVTAPELNSIIRQQL QAHQLSQLQALALPLTPLPVGLQPPSLPAVSAGTGLLSLSALG SQTHLSKEDKNGHDGDTHQEDDGEKSD
Domains • Single domain proteins - • Epidermal growth factor • Serine Proteases - Trypsin • Multi domain proteins -Factor IX -one Ca2+ binding, two EGF/ one protease domain. • Permit building of novel functions by swapping of domains
Factor IX Domain Structure Ca EGF EGF CT Ca - Calcium binding domain EGF - Epidermal growth factor domain CT - Chymotrypsin domain
Chou - Fasman Prediction of Secondary Structure • Based upon analysis of known structures (1974). • Frequency of occurrence of each amino acid in: • a helix • b strand • turn
Chou - Fasman Prediction • List is then analyzed for stretches of amino acids that have a common tendency to form a given secondary structure. • Extend until a region of high probability for either a turn or region with a low probability of both a or b is encountered. • Window is typically <10