Supplementary Figure 1
Myc. Supplementary Figure 1. + FLAG-TWEAK. A. NT 30 min 30 min. a nti-FLAG anti-FLAG control IgG.

Supplementary Figure 1
E N D
Presentation Transcript
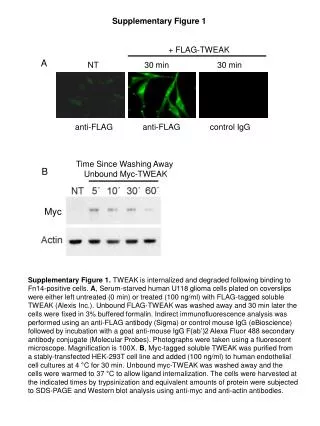
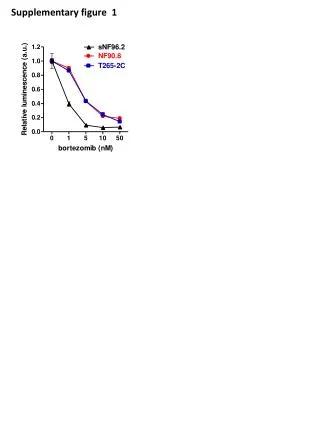
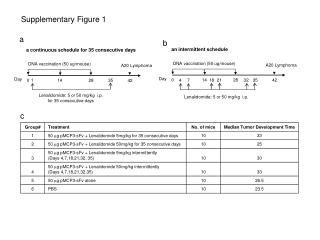
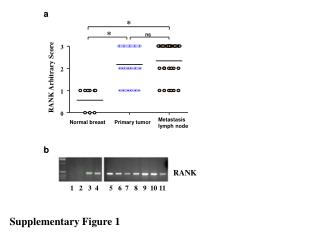
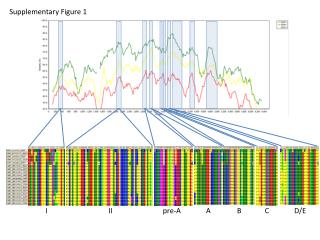
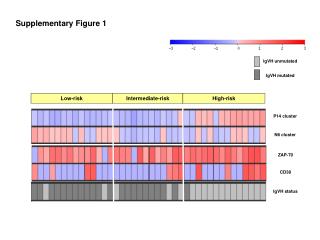
Myc Supplementary Figure 1 + FLAG-TWEAK A NT 30 min 30 min anti-FLAG anti-FLAG control IgG Supplementary Figure 1. TWEAK is internalized and degraded following binding to Fn14-positive cells. A, Serum-starved human U118 glioma cells plated on coverslips were either left untreated (0 min) or treated (100 ng/ml) with FLAG-tagged soluble TWEAK (Alexis Inc.). Unbound FLAG-TWEAK was washed away and 30 min later the cells were fixed in 3% buffered formalin. Indirect immunofluorescence analysis was performed using an anti-FLAG antibody (Sigma) or control mouse IgG (eBioscience) followed by incubation with a goat anti-mouse IgG F(ab’)2 Alexa Fluor 488 secondary antibody conjugate (Molecular Probes). Photographs were taken using a fluorescent microscope. Magnification is 100X. B, Myc-tagged soluble TWEAK was purified from a stably-transfected HEK-293T cell line and added (100 ng/ml) to human endothelial cell cultures at 4 °C for 30 min. Unbound myc-TWEAK was washed away and the cells were warmed to 37 °C to allow ligand internalization. The cells were harvested at the indicated times by trypsinization and equivalent amounts of protein were subjected to SDS-PAGE and Western blot analysis using anti-myc and anti-actin antibodies. Time Since Washing Away Unbound Myc-TWEAK B
Supplementary Figure 2 A GrB-TWEAK 50 nM 25 nM TWEAK 50 nM 12.5 nM Kd=8 nM 25 nM 6.25 nM 12.5 nM 6.25 nM Kd=3 nM B GrB-Fc-IT4 Kd=18 nM 50 nM 25 nM 12.5 nM 6.25 nM Supplementary Figure 2.Surface plasmon resonance analysis of TWEAK, GrB-TWEAK and GrB-Fc-IT4 binding to recombinant Fn14 extracellular domain. Sensogramsshowing binding of TWEAK (A), GrB-TWEAK (A) and GrB-Fc-IT4 (B) to Fn14 immobilized on a CM5 chip. GrB did not bind to immobilized Fn14.
Supplementary Figure 3 GrB-TWEAK, 100 nM, no 1°Ab MDA-MB-231 Non-treated GrB-TWEAK 100 nM GrB-TWEAK 50 nM Supplementary Figure 3.GrB-TWEAK specifically internalizes into Fn14-expressing MDA-MB-231 cells. Cells were untreated or treated with 50 or 100 nmol/L GrB-TWEAK for 5 hours. The cells were fixed, acid washed to remove surface-bound material, permeabilized, and immunostained for the presence of GrB(green). The cells were counterstained with propidium iodide (red) to identify nuclei and visualized using a confocal (Zeiss LSM 510) microscope.
Supplementary Figure 4 Supplementary Figure 4. Effect of GrB-TWEAK in combination with 5-FU, Cisplatin, GemZAR and Doxorubicin on MDA-MB-231 cells. Cells (2,000 cells/well) were seeded in 96-well plates and treated with ~ IC25 doses of GrB-TWEAK (10 nM for MDA-MB-231) and with various concentrations of each chemotherapeutic agent as described in Supplementary Materials and Methods. After 72 h, cell viability was assessed. Normalized isobolograms were then generated using the CalcuSynsoftware, depicting CI values of combination drug studies. CI < 1, CI = 1, and CI > 1 indicate synergism, additive interaction, and antagonism, respectively. No difference in sensitivity to GrB-TWEAK was observed based on the order of treatment. Data shown is from pre-treatment of cells with chemotherapeutic agents prior to addition of GrB-TWEAK. All experiments were performed in triplicate.
Supplementary Figure 5 MDA-MB-231 HT-29 PARP Cleaved PARP Supplementary Figure 5.Caspase-independent PARP cleavage. Pre-incubation of MDA-MB-231 and HT-29 cells with z-VAD-fmk(50 μM) for 1 h prior to 100 nMGrB-TWEAK treatment for 2 hours did not fully block PARP cleavage. β-actin -- + -- + -- + -- + 50 µM z-VAD-fmk -- -- + + -- -- + + 100 nMGrB-TWEAK
Supplementary Figure 6 MDA-MB-231 MDA-MB-435 AAB-527 ME-180 HT-1080 U87-MG SKOV3 Calu-3 HT-29 WM35 PC-3 N-87 T-24 eB1 PI-9 Fn14 β-actin MDA-MB-231 HCC1703 Capan-1 L3.6P1 Capan-2 HCC827 AsPc1 H2073 H1975 A549 HT-29 H358 U87 N87 PI-9 (longer exposure) PI-9 (shorter exposure) Fn14 Supplementary Figure 6. Expression of the GrBinhibitor proteinase inhibitor 9 (PI-9) in various cancer cell lines. Equivalent amounts of whole cell lysate were subjected to SDS-PAGE and Western blot analysis using anti-Fn14, PI-9 and β-actin antibodies. β-actin
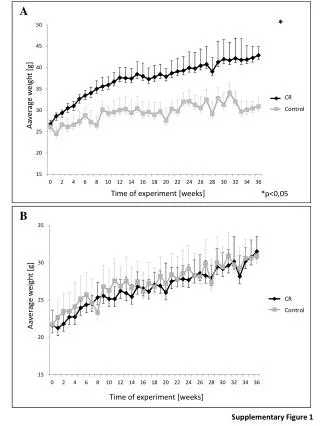
Supplementary Figure 7 A B Supplementary Figure 7. Percent change in body weight of each group of mice from the xenograft experiments, plotted as a function of time. A. GrB or GrB-TWEAK was administered (i.v.) to mice bearing HT-29 xenograft tumors.B.Administration of GrB-TWEAK or GrB-Fc-IT4 into mice bearing MDA-MB-231/Luc breast tumor orthotopicxenografts. Arrows indicate treatment days
Supplementary Figure 8 Supplementary Figure 8. Schematic of GrB-Fc-IT4 fusion protein. GrB-Fc-IT4 is a homodimer in which an Fc domain of human IgG1 is covalently linked to GrB (N-terminus) and the anti-Fn14 humanized scFv of ITEM-4 (C-terminus). Also shown are the disulfide bridge of hinge and the approximate position of the N-linked oligosaccharides attached at Asn297 in the IgG1 Fc-domain.