Protein Structure
330 likes | 357 Vues
Explore the relationship between protein structure and function, from amino acid sequences to tertiary structures. Learn about the peptide bond, constraints on rotation, secondary structures like alpha helices and beta sheets, and stabilizing interactions. Discover the impact of factors like disulfide bonds, prosthetic groups, and coenzymes on protein folding.

Protein Structure
E N D
Presentation Transcript
Protein Structure BL4010 09.26.06
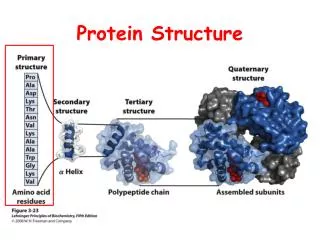
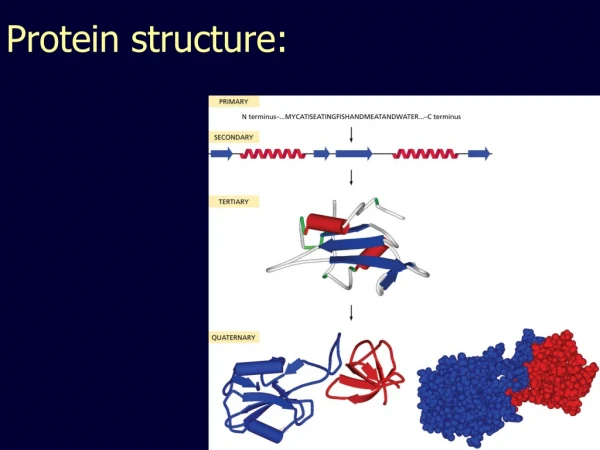
The relationship of structure and function Desirable conformations will be at energy minima 1° structure: amino acid sequence 2° structure: structures localized to certain short stretches of the polypeptide chain - form wherever possible - stabilized by large numbers of H-bonds 3° structure: overall folding of the entire polypeptide 4° structure: overall structure for multimeric proteins (several polypeptides)
The Peptide Bond • 0.133 nm (1.33 Å) - shorter than a typical single bond but longer than a double bond • 40% double bond character • the six atoms of the peptide bond group are planar (C,C=O,N-H, C) • Rotation in the polymer occurs at C • Inherent dipole (N partially positive; O partially negative)
Limited Rotation about Peptide Bond • Two degrees of freedom per residue for the peptide chain • Backbone and side groups limited free rotation
Backbone Torsion Angles • ω angle tends to be planar (0º - cis, or 180 º - trans) due to delocalization of carbonyl pi electrons and nitrogen lone pair • φ and ψ are flexible, therefore rotation occurs here • However, φ and ψ of a given amino acid residue are limited due to steric hindrance • Only 10% of the {φ, ψ} combinations are generally observed for proteins • First noticed by G.N. Ramachandran
Computed Ramachandran Plot Plot of φ vs. ψ The computed angles which are sterically allowed fall on certain regions of plot White = sterically disallowed conformations (atoms come closer than sum of van der Waals radii) Blue = sterically allowed conformations
Experimental Ramachandran Plot X-ray crystallography
Secondary Structure • Repeating values of φ and ψ along the chain result in regular structure • The ability to do this is dependent on steric considerations...i.e. secondary structure is dependent to some degree on primary structure (sequence)
Secondary Structure - alpha helix • For example, repeating values of φ ~ -57° and ψ ~ -47° give a right-handed helical fold (the alpha-helix) e.g. cytochrome c, an alpha helical protein
Secondary Structure - beta sheet Similarly, repetitive values in the region of φ = -110 to –140 and ψ = +110 to +135 give beta sheets. Plastocyanin is composed mostly of beta
Note more allowed regions due to less steric hindrance - Turns
Name φ ψ Structure ------------------- ------- ------- --------------------------------- alpha-L 57 47 left-handed alpha helix 3-10 Helix -49 -26 right-handed. π helix -57 -80 right-handed. Type II helices -79 150 left-handed helices formed by polyglycine and polyproline. Collagen -51 153 right-handed coil formed of three left handed helicies.
Hydrogen Bonding And Secondary Structure beta-sheet alpha-helix
Alpha helix • Residues per turn: 3.6 • Rise per residue: 1.5 Angstroms • Rise per turn (pitch): 3.6 x 1.5A = 5.4 Angstroms • The backbone loop that is closed by any H-bond in an alpha helix contains 13 atoms • phi = -60 degrees, psi = -45 degrees • The non-integral number of residues per turn was a surprise to crystallographers
Beta sheet • Postulated by Pauling and Corey (1951) • Strands may be parallel or antiparallel • Rise per residue: • 3.47 Angstroms for antiparallel strands • 3.25 Angstroms for parallel strands • Each strand of a beta sheet may be pictured as a helix with two residues per turn
Beta turn • allows the peptide chain to reverse direction • carbonyl C of one residue is H-bonded to the amide proton of a residue three residues away • proline and glycine are prevalent in beta turns
Turns & Random Coils • Loops & Turns ( turns) • 1/3 globular protein • Mostly at surface of protein • allows the peptide chain to reverse direction • C=O H-bonded to the NH three residues away • proline and glycine • Random coil • can't assign 2° structure, adopts multiple conformations depending on conditions but not random - energy minima • flexible linkers, hinges
Structure Stabilizing Interactions • Noncovalent • Van der Waals forces (transient, weak electrical attraction of one atom for another) • Hydrophobic (clustering of nonpolar groups) • Hydrogen bonding • Covalent • Disulfide bonds
Disulfide Bonds • Side chain of cysteine contains highly reactive thiol group • Two thiol groups form a disulfide bond • Contribute to the stability of the folded state by linking distant parts of the polypeptide chain
Other factors that affect 2° structure • Prosthetic groups • Coenzymes • Cations • Intramolecular/Intermolecular bonds • disulfides • dityrosine • aldol cross-linking
Tertiary Structure • The backbone links between elements of secondary structure are usually short and direct • Proteins fold to make the most stable structures (make H-bonds and minimize solvent contact
Protein classification • Structural motif • Biochemical function
Protein evolution • Divergent evolution • Similar sequence • Different function • Convergent evolution • Different sequence • Similar function