Molecular Responses to Hypoxia Induced by Submergence in Plants
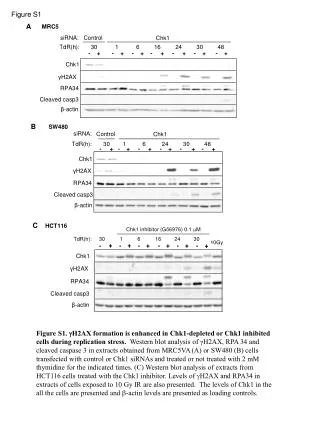
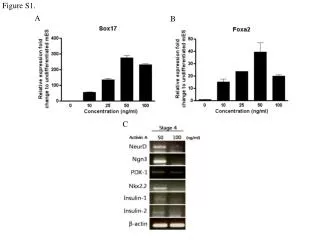
Analysis of CaRLK1 gene expression in response to submergence-induced hypoxia. Ectopic CaRLK1 gene expression affects amino acid biosynthesis genes. Measurements of NtGLB1, NtEF1α, and free amino acids. Western blot analysis of NtRBOHD protein levels.

Molecular Responses to Hypoxia Induced by Submergence in Plants
E N D
Presentation Transcript
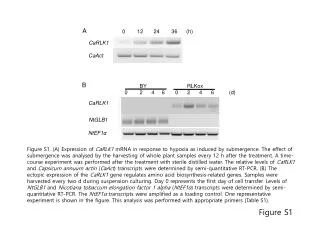
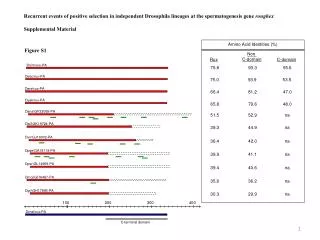
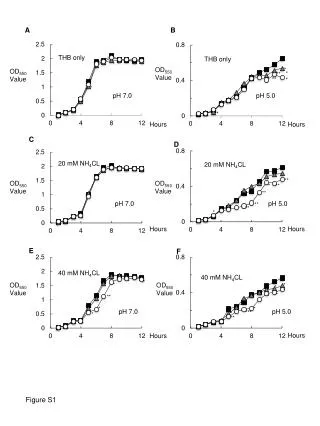
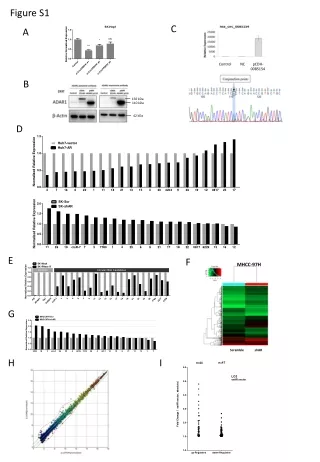
BY RLKox 0 2 4 6 0 2 4 6 (d) A 0 12 24 36 (h) CaRLK1 NtGLB1 CaAct CaRLK1 B Figure S1. (A) Expression of CaRLK1 mRNA in response to hypoxia as induced by submergence. The effect of submergence was analysed by the harvesting of whole plant samples every 12 h after the treatment. A time-course experiment was performed after the treatment with sterile distilled water. The relative levels of CaRLK1 and Capsicum annuum actin (CaAct) transcripts were determined by semi-quantitative RT-PCR. (B) The ectopic expression of the CaRLK1 gene regulates amino acid biosynthesis-related genes. Samples were harvested every two d during suspension culturing. Day 0 represents the first day of cell transfer. Levels of NtGLB1 and Nicotiana tobaccum elongation factor 1 alpha (NtEF1α) transcripts were determined by semi-quantitative RT-PCR. The NtEF1α transcripts were amplified as a loading control. One representative experiment is shown in the figure. This analysis was performed with appropriate primers (Table S1). NtEF1α Figure S1
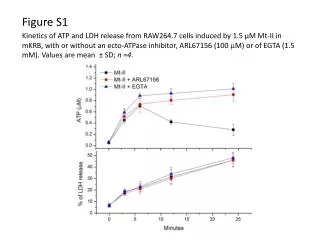
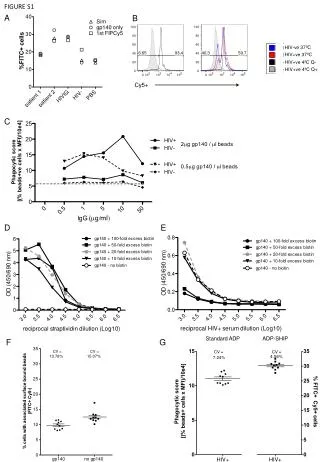
B A Dissolved Oxygen (mg·L-1) Figure S2. (A) Quantification of free amino acids. Each composition was determined using Hitachi’s Amino Acid Analyzer. Error bars represent the standard deviation from the mean (n = 2). (B) Validation of the hypoxic conditions that were induced by the suspension cell culture (A). Dissolved oxygen concentrations in liquid culture medium were analysed after 7 d of culturing. Error bars represent the standard error from the mean (n = 3). The differences between the three conditions are significant at P<0.05. AA (μmol·g-1 DW) Figure S2
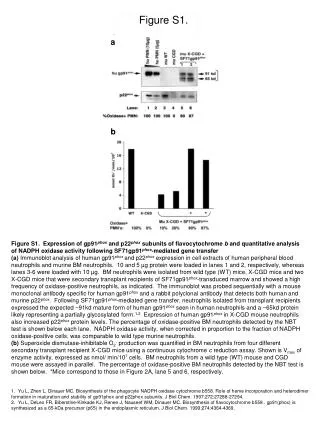
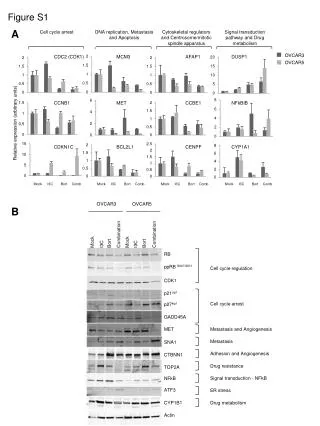
B A 95 43 Figure S3. (A) NtRBOHD protein levels were increased in the RLKox cells. The proteins were extracted, resolved by SDS-PAGE, and analysed by western blot with the anti-NtRBOHD antibody, which was directed against 2 peptides of the NtRBOHD protein, as described in Simon-Plas et al. (2002). A total of 25 μg of protein was loaded. A protein band (43 KDa) that was used as a loading control was detected non-specifically. The molecular mass markers that are indicated on the right sides denote the molecular masses in kDa. (B) Detection of hydrogen peroxide by DAB staining in N. tabacum-originated cells. Red bar indicates length. 200 μm NtRBOHD BY RLKox 43 KDa Figure S3 BY RLKox
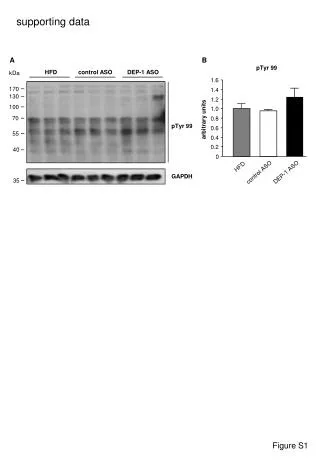
WT RLKox 3 Figure S4. Recovery phenotype of wild-type N. benthamiana and transgenic plants. Four-week-old seedlings were flooded for 8 d and photographed on d 25 after the termination of the flood treatment. Figure S4