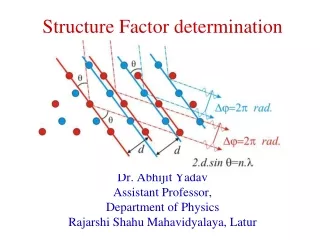
Structure Determination
Structure Determination. High resolution techniques - EM, NMR, X-ray Biochemical techniques -


Structure Determination
E N D
Presentation Transcript
Structure Determination High resolution techniques - EM, NMR, X-ray Biochemical techniques - 1. Use chemicals that react with DNA so that cleavage occurs at all bases (not sequence selective) so as to create a DNA cleavage ladder; alterations in uniform cleavage ladder can be used to footprint DNA 2. Basic strategy to probe DNA conformation with chemicals is to generate some type of conformation-related DNA modification that can be converted into a strand break which can then be mapped Reagents react with one specific base Specificity with respect to: specific base specific base when it is in a unique conformation chemical not specific but method used to generate strand break is
Structure Determination Hydrolysis reaction: Used by nucleases (ribo- and deoxy-) Used by catalytic RNAs
Structure Determination Nucleases
Structure Determination Nucleases -restriction endonucleases Sequence-specific 1. Compatible cohesive ends EcoRI restriction endonuclease GAATTC CTTAAG G AATTC CTTAA G 2. Blunt ends HaeIII restriction endonuclease GGCC CCGG GG CC CC GG
Structure Determination Primary Maxam-Gilbert chemical sequencing of DNA uses base-specific chemical reactions to break chain at A, G, C or T G + A piperidine in acid C + T hydrazine and piperidine in base G DMS and piperidine in base C hydrazine, 1.5 M NaCl and piperidine in base 1. A base-specific rxn either cleaves aromatic ring of base or breaks glycosidic bond 2. The first step destabilizes deoxyribose ring and allows attack by piperidine with subsequent ring cleavage 3. Cleavage of phosphodiester chain 4. Conversion of phosphodiester to phosphomonoester
Structure Determination Primary Maxam-Gilbert chemical sequencing of DNA G + A piperidine in acid
Structure Determination Primary Maxam-Gilbert chemical sequencing of DNA G DMS and piperidine in base
Structure Determination Primary Maxam-Gilbert chemical sequencing of DNA C + T hydrazine and piperidine in base C hydrazine, 1.5 M NaCl and piperidine in base
Structure Determination - Primary Sanger dideoxy sequencing of DNA 5’-AATCGACTAGTACTAGTCTAGCTA-3’ 3’-TTAGCTGATCATGATCAGATCGAT-5’ Denature, 95 ˚C 5’-AATCGACTAGTACTAGTCTAGCTA-3’ + 3’-TTAGCTGATCATGATCAGATCGAT-5’ Anneal radiolabeled primer **5’-AATCGACTAGT 3’-TTAGCTGATCATGATCAGATCGAT-5’ Extend with DNA polymerase dCTP, dATP, dGTP, dTTP ddCTP ddTTP ddATP ddGTP **5’-AATCGACTAGTAddC **5’-AATCGACTAGTACddT **5’-AATCGACTAGTACTAGTddC **5’-AATCGACTAGTACTAGddT **5’-AATCGACTAGTACTAGTCTAGddC **5’-AATCGACTAGTACTAGTCddT **5’-AATCGACTAGTACTAGTCTAGCddT **5’-AATCGACTAGTACTAddG **5’-AATCGACTAGTddA **5’-AATCGACTAGTACTAGTCTAddG **5’-AATCGACTAGTACTddA **5’-AATCGACTAGTACTAGTCTddA **5’-AATCGACTAGTACTAGTCTAGCTddA Denature Gel electrophoresis
Structure Determination - Primary Sanger dideoxy sequencing of DNA 2’ 3’
Structure Determination - Primary Sanger dideoxy sequencing of DNA
Structure Determination - Primary Determination of RNA sequence 1. First convert RNA to DNA using reverse transcriptase 2. Peattie Chemical sequencing of RNA (similar to M-G) uses base-specific chemical reactions to break chain at A, G, C or U Uses aniline at pH 4.6 to cleave RNA at site of modification G DMS and aniline U hydrazine and aniline A > G DEPC and aniline C > U hydrazine and 3 M NaCl and aniline 3. Ribonuclease sequencing RNases are base-specific, cut on the 3’ side of nucleotide label RNA at 5’ or 3’ end, add RNase, PAGE, also run OH- hydrolysis ladder
Structure Determination - Primary 2. Peattie Chemical sequencing of RNA (similar to M-G) G DMS and aniline U hydrazine and aniline A > G DEPC and aniline C > U hydrazine and 3 M NaCl and aniline
Structure Determination - Secondary Probe reactive sites with chemicals - if ds or ss only some sites can be modified For RNA and DNA covalent rxns can reveal which bases are single-stranded (unbonded and usually most reactive), which are in base-pairs (hydrogen bonded) and which are maybe involved in 3˚ interactions Precise locations of binding of proteins & drug molecules can be determined by differences in reactivity of bound and naked DNA Experiments: Radiolabel DNA or RNA At 5’-end with [-32P] ATP and polynucleotide kinase At 3’-end with RNA ligase and radioactive pNp Chemical reactivity determined by: 1. Chemical cleavage of an end-labeled chain at site of modification refer to M-G seq and use of aniline (RNA) and piperidine (DNA) 2. Chain termination by reverse transcriptase at this site RT makes a complementary copy of RNA or DNA single strands, but it stops or pauses at modified bases or at the end of the strand where cleavage occurred. Length of copy identifies location of modified base (seq rxns done in parallel to provide exact position)
Structure Determination - Secondary Probe reactive sites with chemicals
Structure Determination - Secondary Probe reactive sites with chemicals All nitrogens and oxygens in base, sugar & phosphate can be alkylated Rxns with A and C << in dsDNA
Structure Determination - Secondary Probe reactive sites with chemicals - all these monitor W-C base pairing ability Good probes of duplex formation
Structure Determination - Secondary Probe reactive sites with chemicals - react faster in ss regions
Structure Determination - Secondary RNA - use all reagents in Table 3-2 Base specific nucleases (Table 3-1) are all more reactive on ssRNA than on dsRNA S1 hydrolyzes ssRNA or ssDNA but it is inhibited by ds structure Ex: in tRNA only anticodon loop is cut RNase V1 is specific for dsRNA and stacked ssRNA LIMITS of Nucleases - size of enzymes limit accessibility to site OH- cleavage nonspecific cleavage of backbone
Structure Determination - Secondary DNA - mostly duplex but do get ssDNA at cruciforms, triplex, junctions Can also use chemical reagents to determine where proteins bind - FOOTPRINT Cruciforms - ss loops hyperactive to ss-selective reagents bromoacetaldehyde, kethoxal, OsO4, bisulfite Triplex - induced in polypurine rich regions by (-) supercoiling and low pH, in H-DNA ss released and becomes hyperactive to ss-selective reagents, also G in triplex is protected from alkylation at N7 by DMS Nucleases - S1 hydrolyzes ssDNA/RNA FeII(EDTA) is an effective cleaving agent because hydroxyl radicals are oxidative agents that form when ferrous ions are in the presence of O2 (H2O2) and a reducing agent (ascorbate) •OH cleaves the sugar-phosphate backbone of DNA FeII(EDTA) complexes have been applied to DNA and to probing the solvent accessibility of RNA
Structure Determination - Secondary DNA Fe(EDTA)2- + H2O2 + ascorbate Fe2+ H2O2 •OH •OH •OH OH- •OH •OH Fe3+ O P O- O Base O O •OH O P O- O O O- O P O- O Base falls off P O- O O O-
Structure Determination - Secondary DNA Methidiumpropyl-EDTA + Fe (II) + O2 This reagent binds via intercalation, with little sequence discrimination and delivers oxidizing agent to the DNA
Structure Determination - Secondary NAIM - Nucleotide Analog Interference Mapping We’ll discuss a paper
Structure Determination - Tertiary 1. use all chemicals/enzymes used for secondary structure because tertiary contacts can protect regions from chemical modification or nuclease cleavage 2. Chemical crosslinking - using psoralen and/or UV light reagents have 2 reactive groups that can be used to covalently link nucleotides that are far apart in sequence; determine location by EM or partial hydrolysis Combine crosslinking data with chemical probing and phylogeny to get a precise 3-dimensional structure
Structure Determination - Tertiary Chemical crosslinking - using psoralen Used with 16S rRNA and viral DNA 1. Intercalation of psoralen (intercalators are planar and stack as bases) 2. Psoralen absorbs a photon ( = 300-400 nm) 3. C=C on psoralen reacts by cycloaddition to C5=C6 of pyrimidine 4. Psoralen can then form a second crosslink to another pyrimidine T, U >> C
Structure Determination - Tertiary Chemical crosslinking - using UV light Use 200-300 nm light to cause crosslinks in DNA Formation of pyrimidine dimers linked by a cyclobutyl ring across C5=C6 Advantage because no chemical agent has to be added to solution
Structure Determination - Secondary Intercalation - planar aromatic molecules binding to DNA
Structure Determination - Secondary Intercalation - separation of base pairs with a lengthening of double helix and a decrease in helical twist (unwinding)
Structure Determination - Secondary Intercalator - ditercalinium is a bisintercalator (rigid chain) Used as an anticancer drug because it induces DNA repair in many spots on the cell’s DNA (nonspecific, noncovalent binding) which eventually causes the cell to die because of extensive repair-induced DNA degradation