Hidden Markov Models for CpG Island Detection at Tamkang University
Explore methods using Hidden Markov Models to identify CpG islands in genomic sequences. Learn about state path probabilities, Viterbi algorithm, and calculation of posterior state probabilities.

Hidden Markov Models for CpG Island Detection at Tamkang University
E N D
Presentation Transcript
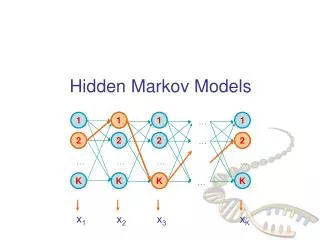
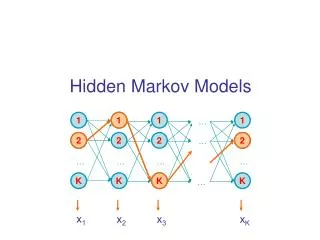
Hidden Markov models Tamkang University Chichang Jou
Questions • Given a short stretch of genomic sequence, how to decide whether it is from a CpG island? • Section 3.1 • Given a long sequence, how do we find the CpG islands in it? • Section 3.2 • Windows of 100 unsatisfactory if CpG islands have sharp boundaries and are of variable lengths
Questions • Given an HMM and a sequence, what is the most • probable state path • Given an HMM, how likely is a sequence, • what is the overall probability of all the probable • path for this sequence • Given an HMM and a sequence, what is the most • probable state for the i-th position of the sequence • (posterior state probability) • Given an HMM without probability parameters and a • set of sequences, how to estimate the parameters
The value may be very small. Veterbi is normally done in log space
Posterior State Probabilities: by backward algorithms • First calculate the probability of producing the sequence with the i-th symbol • in state k bk(i) fk(i)