Phylogenetic Analysis of Rhizobium vitis Strains via Neighbor-Joining Method
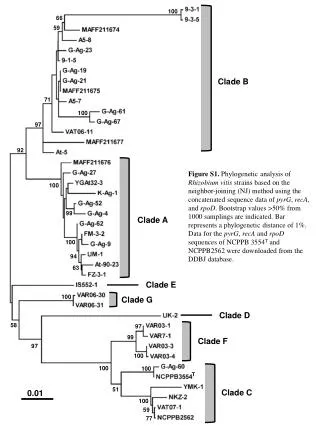
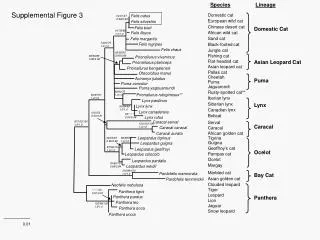
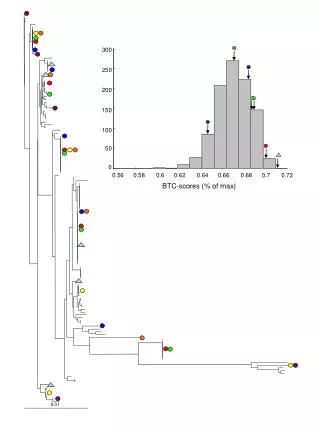
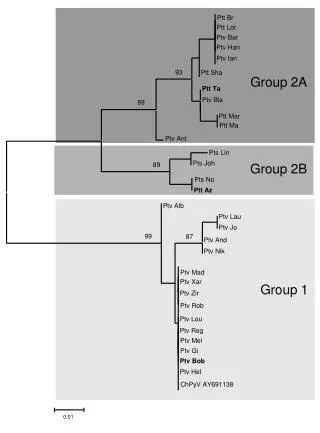
This study investigates the phylogenetic relationships among various strains of Rhizobium vitis using the neighbor-joining (NJ) method. The analysis is based on concatenated sequence data from three genetic markers: pyrG, recA, and rpoD. Bootstrapping with 1000 samplings yielded values greater than 50%, providing robust statistical support for the inferred phylogenetic tree. A phylogenetic distance bar indicates a 1% distance. Sequence data for strains NCPPB 3554T and NCPPB 2562 were sourced from the DDBJ database. The results reveal distinct clades among the studied strains.

Phylogenetic Analysis of Rhizobium vitis Strains via Neighbor-Joining Method
E N D
Presentation Transcript
100 66 59 Clade B 71 100 97 92 Figure S1. Phylogenetic analysis of Rhizobium vitis strains based on the neighbor-joining (NJ) method using the concatenated sequence data of pyrG, recA, and rpoD. Bootstrap values >50% from 1000 samplings are indicated. Bar represents a phylogenetic distance of 1%. Data for the pyrG, recA and rpoD sequences of NCPPB 3554T and NCPPB2562 were downloaded from the DDBJ database. 100 99 Clade A 100 94 63 Clade E 100 Clade G Clade D 58 97 99 Clade F 97 100 100 100 T 51 Clade C 0.01 100 59 77