Face Recognition Using Face Unit Radial Basis Function Networks
Face Recognition Using Face Unit Radial Basis Function Networks. Ben S. Feinstein Harvey Mudd College December 1999. Original Project Proposal. Try to reproduce published results for RBF neural nets performing face-recognition. Recap of RBF Networks.

Face Recognition Using Face Unit Radial Basis Function Networks
E N D
Presentation Transcript
Face Recognition Using Face Unit Radial Basis Function Networks Ben S. Feinstein Harvey Mudd College December 1999
Original Project Proposal • Try to reproduce published results for RBF neural nets performing face-recognition.
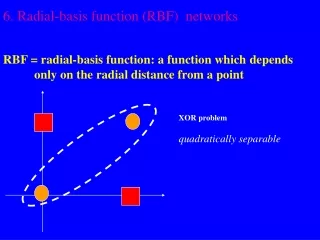
Recap of RBF Networks • Neuron responses are “locally-tuned” or “selective” for some range of input space. • Biologically plausible: Cochlear stereocilia cells in human ear exhibit locally-tuned response to frequency. • Contains 1 hidden layer of radial neurons, usually gaussian functions. Hidden layer output fed to output layer of linear neurons.
Face Unit Network Architecture • First proposed in June 1995 by Dr. A. J. Howell, School of Cognitive and Computing Sciences, Univ. of Sussex, UK. • A face unit is structured to recognize only one person, using hybrid RBF architecture. • Network has two linear outputs, one indicating a positive ID of the person, the other a negative ID.
Face Unit Architecture (2) • An p+a face unit network has p radial neurons linked to the + output, and a neurons linked to the - output. • Challenges • Bitmap faces are big dimensionally • How to reduce dimensionality of problem, extracting only the relevant information?
Gabor Wavelet Analysis • Answer: Use 2D Gabor wavelets, class of orientation and position selective functions. • In this case, reduces dim from |10,000| (100x100 pixel sample) to |126|. • Biologically plausible: Cells in visual cortex respond selectively to stimulation that is both local in retinal position and local in angle of orientation.
Approach to Problem • Sample data • 10 people x 10 poses of each person ranging from 0° (head-on) to 90° (side profile) = 100 sample images • All images 384x287 pixel grayscale Sun rasterfiles, courtesy of Univ. of Sussex face database. • 5 men and 5 women in sample set, mostly Caucasian.
Approach to Problem (2) • Example of images for 1 person...
Approach to Problem (3) • Preprocessing • Used a 100x100 pixel window around pixel at tip of the nose. • Wrote NosePicker Java app to display images and save manually clicked nose coordinates. • Used Gabor orientations (0°, 60°, 120°) with sine and cosine masks = 6 functions. • Calculated the 6 Gabor masks on 99x99, 4 51x51, and 16 25x25 pixel subsamples = |126|.
Approach to Problem (4) • Preprocessing • Sampling windows and orientations...
Approach to Problem (5) • Network Setup/Training • All input vectors were unit normalized, and the unit normalized gaussian function was used. • For each p+a face unit network, fixed set of p poses were used to center the + neurons. • For each + neuron, the nearest p/a unique negative input vectors are used to center p/a - neurons.
Approach to Problem (6) • Network Setup/Training, Cont. • Setting appropriate widths for + and - neurons remains a problem. • Linear output weights are computed by finding the pseudoinverse of the matrix of hidden neuron outputs for each input, A. • Since we want Aw = d => w = A-1d • Used singular value decomposition method to approximate A-1 since A is singular.
Approach to Problem (7) • Network Setup/Training, Cont. • Advantages are instantaneous “training”, since training is no longer iterative process, unlike gradient descent. • Only need to find pseudoinverse and perform matrix vector multiplication to calculate linear output weight vector.
Results • Currently have tested 3+6 and 6+12 networks. • Selection of neuron widths remains a problem, with manual tweaking necessary for good results. • 3+6 performs about like a random classifier.
Results (2) • 6+12 network performed better (see below) • Min correct Min pro Min anti • 37.8% 0 37.2% • Max correct Max pro Max anti • 95.1% 100% 98.7% • Avg correct Avg. pro Avg. ant • 72.6% 55.0% 73.5%
Results (3) • Compare with Dr. Howell (see below) • Avg correct Min pro Min anti • 89% 50% 83 • Max pro Max anti • 100% 100% • Better, however Dr. Howell used a more complex preprocessing scheme, yielding input vectors of |510|.
Future Work • Devise algorithm to choose appropriate neuron widths for + and - neurons or experiment with other radial basis functions that don’t need widths, such as the thin spline. • Implement a network of face units, whose output will indicate a face’s identity instead of just an affirmative or negative response.
Future Work (2) • Implement a confidence threshold to automatically discard low-confidence results. • Expand Gabor preprocessing scheme to yield more coefficients.
What Code Was Written? • Wrote C++ RBFNet class and rbf app to implement RBF net with n dimensional input and 1 linear output neuron. • Uses k-means clustering, global first nearest neighbor heuristic, and gradient descent. • Wrote C++ FaceUnit class and face_net app to implement a scalable face unit network.
What Code Was Written? (2) • Wrote Java app to display images and save manually clicked nose coordinates. • Wrote C++ program to perform image sampling and Gabor wavelet preprocessing. • Wrote perl scripts to generate input files. Hope to soon have perl script to automatically run input files and compile performance results.
Acknowledgments • Dr. A. J. Howell, School of Cognitive and Computing Sciences, Univ. of Sussex, UK. • Provided Gabor data and sample face images. • Dr. Robert Oostenveld, Dept. of Medical Physics and Clinical Neurophysiology, University Nijmegen, The Netherlands. • Provided C routine for SVD pseudoinverse calculation.
Acknowledgments (2) • Numerical Recipies Software, Numerical Recipies in C: The Art of Scientific Computing. • Used their published singular value decomposition routine in C. • And last, but not least… Prof. Keller • Invaluable guidance and advice regarding this project.