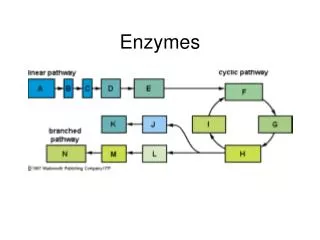
Lecture 5:Enzymes
Lecture 5:Enzymes. Ahmad Razali Ishak Department of Environmental Health Faculty of Health Sciences UiTM Puncak Alam. Introduction to Enzymes. Enzymes are biological catalysts. Like all catalysts, enzymes lower the energy needed to get a reaction started. Enzymes are much


Lecture 5:Enzymes
E N D
Presentation Transcript
Lecture 5:Enzymes Ahmad Razali Ishak Department of Environmental Health Faculty of Health Sciences UiTM Puncak Alam
Introduction to Enzymes Enzymes are biological catalysts. Like all catalysts, enzymes lower the energy needed to get a reaction started. Enzymes are much generally better at accelerating the rates of reactions than non-biological catalysts. Diagram showing that less energy is required to get an enzyme catalyzed reaction started as compared to a non-catalyzed reaction
Enzyme Catalysis (Cont’d) • Consider the reaction
These six classes are: 1. Oxidoreductases - enzymes catalyzing oxidation reduction reactions. 2. Transferases - enzymes catalyzing transfer of functional groups. 3. Hydrolases - enzymes catalyzing hydrolysis reactions. 4. Lyases - enzymes catalyzing group elimination reactions to form double bonds. 5. Isomerases - enzymes catalyzing isomerizations (bond rearrangements). 6. Ligases - enzymes catalyzing bond formation reactions couples with ATP hydrolysis
The Active Site of the Enzyme. • Each enzyme has a unique active site. • Active site = catalytic site. • The enzyme binds its substrate(s) at the active site and the enzyme catalyzes chemical changes in the substrate(s).
Enzymes contain a large number of amino acids, but most AA side chains are used for forming the enzyme's shape. Only a few AA side chains are at the active site. These special AA side chains: 1. Bind the substrate(s) and 2. Catalyze the reaction
3-D model of the enzyme ribonuclease with the key amino acid side chains at the active site shown in red. The active site is a deep groove at the center of this structure. · Enzyme has large structure with hundreds of AA side chains but only a few are involved in catalysis. · Each enzyme has a unique active site. · Key AA side chains are involved in binding and catalysis in the active site.
Enzyme + Substrate = Enzyme-Substrate Complex • E + S = ES Complex
Binding Models • Two models have been developed to describe formation of the enzyme-substrate complex • Lock-and-key model: substrate binds to that portion of the enzyme with a complementary shape • Induced fit model: binding of the substrate induces a change in the conformation of the enzyme that results in a complementary fit
Enzyme Kinetics Usefulness of enzyme kinetics: · Common clinical assays to detect enzymes · Understanding metabolic pathways · Measuring binding of substrates and inhibitors to the active site of an enzyme · Understanding the mechanism of catalysis of an enzyme
A Simple Mechanism for the Enzyme Catalyzed Reaction. • For catalysis to begin, the substrate must bind to the enzyme, which results in the formation of the enzyme-substrate complex (ie E-S complex). • The E-S complex forms rapidly in the first part of the enzyme catalysis process and the concentration of the E-S stays constant at a steady-state level. For this reason, this type of kinetics is called steady-state kinetics.
E + S ↔ ES → E + P These data show that at low [S], the initial velocity is more or less proportional to the [S]. At high [S], the initial velocity no longer increases as more substrate is added. Thus, at high [S] the enzyme is saturated with substrate and no increase in the enzyme catalyzed rate is observed.
The Michaelis-Menten Equation. • The plot of Vo versus [S] can be represented by an equation, which is known as the Michaelis-Menten equation in honor of the scientist who first described it. This equation, sometimes called the M-M equation, is an important one for you to know and understand. Vo = Vmax [S] /Km + [S]
The constants in this equation, Km and Vmax, are defined: Vmax = Maximum velocity catalyzed by a fixed [E] Km = the [S] which gives 1/2 Vmax
The Km is sometimes called the Michaelis Constant. The Km is an intrinsic property of an enzyme related to the binding constant for forming the ES complex, which is an equilibrium and can be defined by the rate constants for its formation and breakdown using the simple enzyme
To calculate the Km and Vmax, the Michaelis-Menten equation is converted into a linear form by taking the reciprocal of both sides of the equation. This is called the Lineweaver-Burk equation in honor of the first scientists to describe it.
This equation then takes on the form of the equation of a line. The y values are 1/Vo, the x values are 1/[S]. The b value in the line equation is the slope and equal to Km/Vmax, while the c value is the y-intercept and equal to 1/Vmax.
Enzyme Inhibitors • Inhibitors of enzymes: Two types are considered - Competitive and Non-Competitive. • Competitive Inhibitor has a chemical similarity to the substrate and competes with the substrate for binding to the active site of the enzyme. A good example to describe competitive inhibition is the mitochondrial enzyme, succinate dehydrogenase:
The reaction catalyzed by succinate • dehydrogenase is the oxidation of succinate • to fumarate. (B) Malonate and oxaloacetate • are competitive inhibitors of succinate • dehydrogenase.
Both these competitive inhibitors, malonate and oxaloacetate, look like succinate in their chemical character. Both inhibitors are dicarboxylic acids like the substrate succinate so they have groups which can bind in the same places in the active site of succinate dehydrogenase as the substrate. • However, neither inhibitor has the capacity to undergo the reaction and so the enzyme is inhibited. • Since these inhibitors simply bind to the enzyme, when the succinate concentration is high, they will be pushed out of the site by the substrate and the enzyme will catalyze the reaction as if no inhibitor were present.
An enzyme mechanism model of the action of a competitive inhibitor (Ic) based on the standard model of a Michaelis-Menten enzyme where E + S leads to the E-S complex, which leads to product P:
Vo versus [S] plot comparing the kinetics of the reaction in the absence of inhibitor and in the presence of the competitive inhibitor (Ic). At high [S], the initial velocity in the presence of Ic will be about the same as it is in the absence of the inhibitor. The concentration of S which will be required to overcome the effect of the competitive inhibitor will depend on the [Ic] (ie. concentration of the competitive inhibitor) and the Ki (ie. the binding constant of the inhibitor to enzyme).
Here the uninhibited reaction gives the standard double reciprocal plot from which Km and Vmax can be calculated. The reaction in the presence of the competitive inhibitor yields apparent constants for the enzyme which are called the Km' and Vmax'. For the true competitive inhibitor, the Vmax' (apparent Vmax for inhibited enzyme) will be the same as the real Vmax, while the Km' (apparent Km for the inhibited enzyme) will be greater than the real Km. Thus, the -1/Km' will be smaller than -1/Km.
Non-competitive Inhibition. • Non-Competitive Inhibitor does not compete with substrate and the [S] has no influence on the degree of inhibition of the enzyme's catalytic rate. For example, enzymes with a thiol ( -SH ) not at the active site can be inhibited
Example of a heavy metal inhibiting an enzyme by binding to a thiol group not at the active site and inactivating the enzyme.
In this case where the non-competitive inhibitor (Inc) reacts with the enzyme at a site other than the active site, both the free enzyme (E) and the enzyme-substrate complex (E-S) react with Inc. Clearly, in this case the reaction of the non-competitive inhibitor is irreversible.
A Vo versus [S] plot for the Non-competitive Inhibitor looks very different than that for a competitive inhibitor since increasing the [S] has no impact
Non competitive inhibitors decrease Vmax but have no effect on Km.
Evaluating Enzyme Inhibitors to determine type and their Ki. To determine what type an inhibitor is: 1. Find Km and Vmax for uninhibited from 1/Vo vs 1/[S] plot. 2. On same graph find Km' and Vmax' for inhibited reaction. A. If Vmax = Vmax' then inhibitor is competitive type. (Vmax and Vmax' should not be more than 10% different) B. If Vmax does not equal Vmax', then if Km = Km', inhibitor is non competitive type.
Assignment 1.PLOT Vo versus [S] to show that the enzyme obeys Michaelis-Menten Equation. 2.PLOT 1/Vo versus 1/[S] to determine Km and Vmax.
1) Draw "vo vs. [S]" and "1/vo vs 1/[S]" plots 2) Determine the Km and Vmax from the double reciprocal plot 3) Decide what type of inhibitors "x" and "z" are 4) Calculate the Ki for "x" and "z" Competitive Inhibitor : Km' = Km (1 + [I]/Ki) Noncompetitive Inhibitor: Vmax' = Vmax / (1 + [I]/Ki)