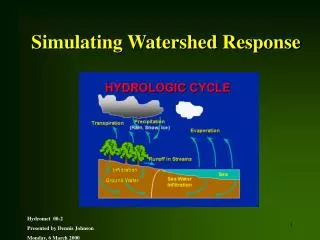
DNA Assembly with Gaps: Simulating Sequence Evolution
DNA Assembly with Gaps: Simulating Sequence Evolution. Reed A. Cartwright Department of Genetics University of Georgia. Synopsis. Explain the importance of simulations. Introduce Dawg, a new sequence simulation program. Example usage of Dawg. Why Simulate Phylogenies?.

DNA Assembly with Gaps: Simulating Sequence Evolution
E N D
Presentation Transcript
DNA Assembly with Gaps: Simulating Sequence Evolution Reed A. Cartwright Department of Genetics University of Georgia
Synopsis • Explain the importance of simulations. • Introduce Dawg, a new sequence simulation program. • Example usage of Dawg. RA Cartwright rac@uga.edu - http://scit.us/
Why Simulate Phylogenies? • Biologists use many techniques to reconstruct phylogenies based on biological data. • However, true phylogenies are unknown, except for a few instances. • How then can we test the accuracy of these reconstruction methods? • Use simulations. RA Cartwright rac@uga.edu - http://scit.us/
Why Simulate Phylogenies? • Techniques are often based on certain models of evolution. • Simulating sequence evolution based on these models produces an ideal situation to test the techniques. • Using other models can test how robust a technique is. RA Cartwright rac@uga.edu - http://scit.us/
A A B B C C D D A B C D Testing Procedure 1. Start with a “known” tree. 3. Estimate the trees of the simulated data. 2. Simulate sequence sets based on the tree. 4. Compare estimated trees to the original tree. A AATTCTTTGAGTTAA B AATTCTTTGAGTTAA C AATTCTTAAAGTTAA D AATTCTTAAAGTTAA A AAAAGATAAAGCAAA--A B GAAAGATAAAGCAAA--A C GAAAGATAAAGAAAAACA D GAAAGATAAAGAAAAACA RA Cartwright rac@uga.edu - http://scit.us/
Simulating Evolution • Proper simulation of molecular evolution should include both substitutions and indels. • However, existing programs either do not include indels or use an unjustified model of indel formation. • Dawg was created to address this gap. RA Cartwright rac@uga.edu - http://scit.us/
What is Dawg? • Dawg stands for “DNA Assembly with Gaps.” • A portable and robust program for simulating molecular evolution. • Development Website: http://scit.us/dawg/ RA Cartwright rac@uga.edu - http://scit.us/
Comparing Software RA Cartwright rac@uga.edu - http://scit.us/
Parameters • Tree phylogeny • TreeScale coefficient to scale branch lengths by • Sequence root sequences • Length length of generated root sequences • Rates rate of evolution of each root nucleotide • Model model of evolution: GTR|JC|K2P|K3P|HKY|F81|F84|TN • Freqs nucleotide (ACGT) frequencies • Params parameters for the model of evolution • Width block width for indels and recombination • Scale block position scales • Gamma coefficients of variance for rate heterogeneity • Alpha shape parameters • Iota proportions of invariant sites • GapModel models of indel formation: NB|PL|US • Lambda rates of indel formation • GapParams parameter for the indel model • Reps number of data sets to output • File output file • Format output format: Fasta|Nexus|Phylip|Clustal • GapSingleChar output gaps as a single character • GapPlus distinguish insertions from deletions in alignment • LowerCase output sequences in lowercase • Translate translate outputed sequences to amino acids • NexusCode text or file to include between datasets in Nexus format • Seed PRNG seed (integers) RA Cartwright rac@uga.edu - http://scit.us/
Sample Input File # example.dawg Tree = ((AY727331:0.001359,AY727330:0.001359):0.084512, (AY727327:0.006116,AY727326:0.006116):0.079756); Model = "GTR" Params = {1.08031, 2.45581, 0.44452, 1.09145, 4.06519, 1.00000} Freqs = {0.353470, 0.143681, 0.178206, 0.324643} Length = 300 Lambda = 0.143120 GapModel = "NB" GapParams = {1, 0.753247} Format = "Clustal" File = "example.aln" Seed = 1981 RA Cartwright rac@uga.edu - http://scit.us/
CLUSTAL multiple sequence alignment (Created by DAWG Version 1.0.0) AY727326 TTCGAAAATATGTTAGTACTCAATATGAATTCTTTGAGTTAAAAAAGATAAAGCAAA--A AY727327 TTCGAAAATATGTTAGTACTCAATATGAATTCTTTGAGTTAAGAAAGATAAAGCAAA--A AY727330 TTCAAAAATATGCTAGGACTGAATATGAATTCTTAAAGTTAAGAAAGATAAAGAAAAACA AY727331 TTCAAAAATATGCTAGGACTGAATATGAATTCTTAAAGTTAAGAAAGATAAAGAAAAACA AY727326 ATACATAATGTGATTTCAATATTCCAATTACCTAACAATACGGCTATCAATTAAACGATT AY727327 ATACATAATGTGATTTCAATATTCCAATTACCTAACAATACGGCTATCAATTAAACGATT AY727330 GTACATAATGTAAA----TTATTGCAA---------AAAACGGCTAACAATTAGACGATT AY727331 GTACATAATGTAAA----TTATTGCAA---------AAAACGGCTAACAATTAGACGATT AY727326 TTAGGATTACACCGACAAATATTAGGCCGATATGAATTTAACATCATGTTGTATTTAGAT AY727327 TTAGGATTACACCGACAAATATTAGGCCGATATGAATTTACCATCATGTTGTATTTAGAT AY727330 TTAGGATTACGCTGACAAATATTAGGATGATATTAATTTA------TCTTGTATTTAGAT AY727331 TTAGGATTACGCTGACAAATATTAGGATGATATTAATTTA------TCTTGTATTTAGAT AY727326 GCTGTCTTTTATTAACATTCATCATTAAAT-TTGGAACCTTTTGCATTTAAGAAGTACAT AY727327 GCTGTCTTTTATTAACATTCATCATTAAAT-TTGGAACCTTTTGTATTTAAGAAGTACAT AY727330 GCTGTCTTTTATCAACATTCATCACTAGATATTGGAACCTATTGCATCTAAGAAGTACAT AY727331 GCTGTCTTTTATCAACATTCATCACTAGATATTGGAACCTATTGCATCTAAGAAGTACAT AY727326 GTTTAATAGTGTTTAAAA-TATATATGAAATTGATCATAAGGA---TCTATAAATGCGGT AY727327 GTTTAATAGTGTTTATAA-TATATATGAAATTGATCGTAAGGA---TCTATAAATGCAGT AY727330 GTTTAATAGGGTT-AAAACTATATATGAAGTCGATTATAAGGAATTTCTATAAATGTAGC AY727331 GTTTAATAGGGTT-AAAACTATATATGAAGTCGATTATAAGGAATTTCTATAAATGTAGC AY727326 TCTTCAATTTCTTG AY727327 TCTTCAATTTCTTG AY727330 TCTTCAATTTCCTA AY727331 TCTTCAATTTCCTA RA Cartwright rac@uga.edu - http://scit.us/
Estimating Indel Rate • Dawg would be of little benefit if biologists could not estimate parameters of indel formation from real data. • Dawg’s indel model allows such estimation, which is implemented in a Perl script, lambda.pl. RA Cartwright rac@uga.edu - http://scit.us/
Example Usage:Confidence Interval of Indel Rate • I aligned the sequences of chloroplast trnK introns from two Hibiscus and two Prunus species. • Using Paup*, I estimated the phylogeny and substitution parameters. • Using lambda.pl, I estimated the indel formation parameters. RA Cartwright rac@uga.edu - http://scit.us/
Example Usage • From these estimated parameters of evolution, I constructed an input file for Dawg. • From the input file Dawg produced a thousand simulated sequence sets. • The rate of indel formation was estimated for each of the simulated sequences. RA Cartwright rac@uga.edu - http://scit.us/
Results • The estimated rate of indel formation was 0.143120. • Bootstrapping gave a 95% CI of 0.078530 to 0.213560. • Biologically this is 8 to 21 indels per 100 substitutions. RA Cartwright rac@uga.edu - http://scit.us/
Synopsis • Explain the importance of simulations. • Introduce Dawg, a new sequence simulation program. • Example usage of Dawg. RA Cartwright rac@uga.edu - http://scit.us/
Marjorie Asmussen Wyatt Anderson John Avise Jim Hamrick Ron Pulliam Paul Schliekelman Jeff Ross-Ibarra Beth Dakin Douglas Theobald Yong-Kyu Kim Thanks RA Cartwright rac@uga.edu - http://scit.us/