Recombinant DNA Technology
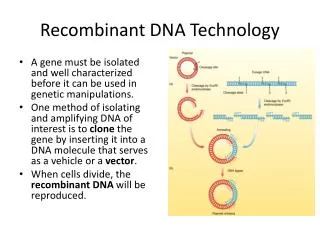
Recombinant DNA Technology. Recombinant DNA Technology. T echnique that allows DNA to be combined from different sources DNA code universal Foundation of genetic engineering. Review. What are enzymes? What function do they serve? What makes up the DNA sequence?


Recombinant DNA Technology
E N D
Presentation Transcript
Recombinant DNA Technology • Technique that allows DNA to be combined from different sources • DNA code universal • Foundation of genetic engineering
Review • What are enzymes? What function do they serve? • What makes up the DNA sequence? • What is the purpose of the genetic code?
Transformation Lab Highlights • Bacterial plasmids • Recombinant DNA plasmids • Transformation of plasmids • Antibiotic selection of recombinant DNA • pGLO plasmid • GFP gene (green fluorescent protein) • Antibiotic (ampicillin) resistant gene
Recombinant DNA Technology http://www.youtube.com/watch?v=x2jUMG2E-ic http://www.youtube.com/watch?v=8rXizmLjegI&feature=related
Discovery of Restriction Enzymes • Bacteriophages are viruses that infect bacteria • Insert their genetic material (DNA) into the DNA of bacteria • Bacterial cells reproduce virus genetic material
Discovery of Restriction Enzymes • In the 1960s, microbiologists discovered some bacteria protected from bacteriophage infection • Because they can restrict phage replication • Restricted growth of phages occurred because the bacteria contained enzymes that could cutviral DNA into small pieces and prevent replication • These enzymes were called restriction enzymes
Restriction Enzymes • Primarily found in bacteria • Given names based on genus and species names of bacteria from which they are isolated • Cut DNA by cleaving the phosphodiester bond (in sugar-phosphate backbone) that joins nucleotides in DNA strand • Enzymes bind to, cut (digest) DNA within specific sequences of bases called recognition sequence or restriction site • Typically recognize 4, 6, or 8 nucleotide sequences
Restriction Enzymes • Restriction enzyme - EcoRI • Named because it was discovered in - Escherichia coli
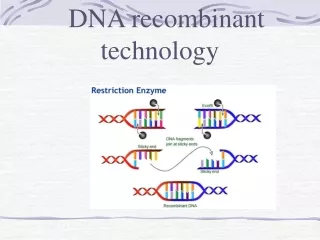
Restriction Enzymes • Some restriction enzymes cut DNA to create DNA fragments with overhanging single-stranded ends called “sticky” or cohesive ends
Restriction Enzymes • Some enzymes generate fragments with non-overhanging or blunt ends
Restriction Fragments • Each cut of DNA creates restriction fragments • A long strand of DNA may be cut several times by a restriction enzyme to create several restriction fragments
Restriction Enzymes • Enzymes that produce cohesive ends are often favored over blunt-end cutters because DNA fragments with cohesive ends can easily be joined together. • DNA from any source, bacteria, humans, dogs, frogs, dinosaurs, and ancient human remains can be digested by a particular restriction enzyme as long as it has the restriction site for that enzyme!
Recombinant DNA History In the early 1970s, a group of scientists joined together DNA from E. coli chromosomes and DNA from a primate virus • They isolated the DNA from each organism and cut them into fragments with EcoRI • They added the E. coli and viral DNA fragments to a reaction tube and added DNA ligase • They created a hybrid of the two DNA types, recombinant DNA
NIH and RAC • Scientists were concerned about: • with what might happen if recombinant bacteria were to leave the lab • if such bacteria could transfer their genes to other cells • survival in other organisms including humans • National Institutes of Health (NIH) formed the Recombinant DNA Advisory Committee (RAC) to evaluate the risks of recombinant DNA technology and establishing guidelines • http://oba.od.nih.gov/rdna_rac/rac_about.html
Recombinant DNA Technology • Plasmids are circular DNA found in bacteria and replicate independently • Plasmidsare considered extrachromosomalDNA because they are present in the bacterial cytoplasm in addition to the bacterial chromosome • Plasmids can be used as vectors, pieces of DNA that can accept, carry, and replicate (clone) other pieces of DNA
Plasmid Vectors – used to clones specific DNA http://www.sumanasinc.com/webcontent/animations/content/plasmidcloning.html
Transformation • Transformation – process for inserting foreign DNA into bacteria • Treat bacterial cells with calcium chloride solutions • Add plasmid DNA to cells chilled on ice • Briefly heat the cell and DNA mixture, plasmid DNA will enter the bacterial cell • Once inside bacteria, plasmids replicate and express their genes
Selection • Joining of DNA fragments and plasmids by transformation can be inefficient • Some plasmids will rejoin and recircularize and not bond with any foreign DNA • During transformation, many cells will not take up DNA • Recombinant bacteria, those transformed with a recombinant plasmid, must be separated from nontransformed bacteria as well as cells with plasmids without foreign DNA • This process is called selection because it is designed to identify (select for) recombinant bacteria and prevent the growth of nontransformed bacteria and bacteria without foreign DNA
Transformation/Selection Animations • http://www.phschool.com/science/biology_place/labbench/lab6/concepts1.html • http://highered.mcgraw-hill.com/sites/0072556781/student_view0/chapter13/animation_quiz_1.html • http://www.dnalc.org/resources/animations/transformation1.html
Transformation/Selection Resources • Academic: • http://faculty.plattsburgh.edu/donald.slish/transformation.html • http://www.genome.ou.edu/protocol_book/protocol_adxF.html • http://www.bio.davidson.edu/Courses/Molbio/MolStudents/spring2003/Siegenthaler/Transformation.htm • Pre-lab Resources: • http://www.escience.ws/b572/L2/L2.htm • http://faculty.clintoncc.suny.edu/faculty/michael.gregory/files/bio%20101/bio%20101%20laboratory/bacterial%20transformation/bacteria.htm
Polymerase Chain Reaction • Technique for making copies or amplifying a specific sequence of DNA in a short period of time.
PCR (Polymerase Chain Reaction) 1. Target DNA to be amplified is added to a thin-walled tube and mixed with deoxyribonucleotides (dATP, dCTP, dGTP, dTTP), buffer and DNA polymerase. Primers are added to the mixture; they are complimentary to nucleotides on opposite ends of the target DNA. 2. Tube is placed in a thermal cycler (gene cycler). This will take the sample through a series of reactions called the PCR cycle.
Thermal Cycler • Denaturation – tube is heated to ~94-96ºC separating DNA into single strands • Hybridization (annealing) – tube is cooled to ~60-65ºC allowing primers to bond to ends of DNA • Extension (elongation) – temperature raised slightly and DNA polymerase copies the DNA by binding to primer and making a new strand DNA polymerase used in this reaction is Taq DNA polymerase from Thermus aquaticcus. Bacteria is adapted to live in hot springs it has enzymes that can withstand high temperatures necessary for PCR.
PCR • At the end of one cycle, amount of DNA has doubled. • Researchers can repeat cycle (usually 20-30 times) to amplify millions of copies of DNA
Uses of PCR • Studying gene expression • Detection of viral or bacterial infections • Diagnosing genetic conditions • Amplifying trace amounts found at crime scenes • Amplifying trace amounts found in ancient DNA http://www.dnalc.org/resources/animations/pcr.html http://www.youtube.com/watch?v=_YgXcJ4n-kQ http://www.maxanim.com/genetics/PCR/pcr.swf http://practicality.wordpress.com/2008/01/13/the-pcr-song-with-lyrics/
Gel Electrophoresis Gel Electrophoresis • Used to separateand visualize DNA fragments based on size • Agarose gel is a semisolid material with small pores through which DNA can travel; fragments are separated by size – smaller fragments can travel further through pores than larger fragments
Gel Electrophoresis Gel Electrophoresis 1. To run a gel, it is submerged in buffer solution that will conduct electricity 2. DNA samples are loaded into small wells and an electric current is passed through the gel 3. The sugar-phosphate backbone renders DNA with a negative charge; therefore DNA will move to the positively charged end • Because migration distance is inversely proportional to the size of the DNA fragment, large DNA fragments migrate short distances and small fragments migrate faster • Dyes are added to monitor DNA migration
Gel Electrophoresis Animations Gel Electrophoresis • http://www.sumanasinc.com/webcontent/animations/content/gelelectrophoresis.html • http://www.dnai.org/b/index.html • http://learn.genetics.utah.edu/units/biotech/gel/ • http://www.youtube.com/watch?v=2UQIoYhOowM
5'GG CC3'3'CC GG5' 5'GGCC3'3'CCGG5' 5'GG CC3'3'CC GG5' Restriction Enzymes • DNA cutting enzymes • Cuts double stranded DNA • Two incisions to cut DNA at sugar-phosphate bond on each strand • Very specific • Enzymes recognize short nucleotide sequences • Enzymes cut the DNA at specific points within DNA • For example – if the restriction enzyme cuts between a C and G then
G A A T T C C T T A A G G A A T T C C T T A A G G AATC CTTAA G Restriction Enzymes • Each restriction enzyme recognizes a specific restriction site, a short nucleotide sequence • Some restriction enzymes cut straight across the DNA • Some restriction enzymes cut at offset nucleotides • For example -
Restriction Enzymes • Sequences that are cut at offset points produce overhanging pieces of DNA called “sticky ends” Sticky end G AATC CTTAA G
Restriction Enzymes http://users.rcn.com/jkimball.ma.ultranet/BiologyPages/R/RestrictionEnzymes.html
Why are restriction fragments useful? • Recombinant DNA • Gene isolation • Genome mapping • Gel electrophoresis • Genetic engineering • Gene therapy