JGI Timeline
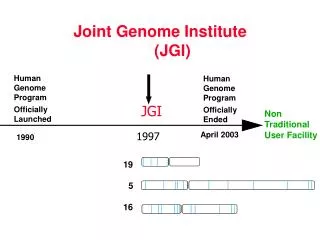
JGI Timeline. Human Genome Program Officially Launched. Human Genome Program Officially Ended. JGI. April 2003. 1990. 1997. 19. 5. 16. Joint Genome Institute …………………. (JGI). Non Traditional User Facility. The JGI Post Human Genome Project Community Sequencing Program


JGI Timeline
E N D
Presentation Transcript
JGI Timeline Human Genome Program Officially Launched Human Genome Program Officially Ended JGI April 2003 1990 1997 19 5 16 Joint Genome Institute ………………….(JGI) Non Traditional User Facility
The JGI Post Human Genome Project Community Sequencing Program (CSP) Microbial Community Genomics
Overview The Community Sequencing Program (CSP) To provide the scientific community through a peer reviewed process access to high throughput sequencing at the JGI.
User Guide > How to Propose a Project What types of projects will the JGI/CSP accept? A wide range of projects. Ultimately, the most important factor in determining if a project will be accepted is its scientific merit.
Proposals & Peer Review Process General Scientific Users Proposals Designated Lab Director Proposal Study Panel Scientific Advisory Committee Users JGI Director Sequence Allocation
What can researchers get from the CSP program? The deliverables can range from raw sequence traces to well-annotated assembled genomes depending on the request in the proposal.
Interactions of the JGI and Scientific Users with Approved Sequencing Proposals Scientific Support for Approved Projects Production Sequencing Users Scientific Support Group SSG Informatic Analysis Of Sequence
Interactions of the JGI and Scientific Users with Approved Sequencing Proposals Scientific Support for Approved Projects Production Sequencing DOE Gov Agencies Scientific Support Group SSG GTL, Microbe CSP (EPA,USDA, NSF) Informatic Analysis Of Sequence
DOE Production Sequencing Informatics JGI Science Programs
DOE+CSP+Gov A Scientific Support Group Informatics Production Sequencing JGI Science Programs
Sequence Based Science at the JGI • Gene Regulatory Vocabulary of Animals • Studies of Body Plan Evolution • Microbial Community Genomics
< 1% of microbes are culturable • Many unculturables live in interdependent consortia of considerable diversity • Aim: to recover genome-scale sequences and reveal metabolic capabilities • What is the structure of natural microbial populations? What is a microbial species? Can we harness their metabolic capabilities
What Enviroments to Study? • Ones with minimal microbial complexity
Iron Mountain JillBanfield Gene Tyson Phil Hugenholtz UC Berkeley Geology Jill Banfield et al. UC Berkeley
Iron Mountain Superfund site Discharging >1 ton of toxic metals/day (pH <1) FeS2
= = = ===== === == = Enviromental Sample Purify High Molecular Weight DNA Fosmid Library Construction Shotgun Library Construction ===== === == = ===== === == = ===== === == = Fosmid Insert End Sequencing = DNA Sequencing Assembly Annotation
Shotgun Library Construction Shotgun Library Construction = = ===== === == = ===== === == = ===== === == = ===== === == = = = = ===== === == = Enviromental Sample Purify High Molecular Weight DNA When possible culture isolates Fosmid Library Construction ===== === == = Fosmid Insert End Sequencing DNA Sequencing Assembly Annotation ? =
Iron Mtn “whole metagenome shotgun” GC content separates into two components bacteria Reverse read average G+C archaea Forward read average G+C
Iron Mountain “whole metagenome shotgun” GC and depth distributions 0.55 Read average G+C 0.38 Read depth Lepto III 3 10 Lepto II Bacterial
0.55 Read average G+C 0.38 Read depth Lepto III 3 10 Lepto II Bacterial Archaeal Fer 1 (cultured and sequenced ) G-plasma Fer 2 3 10
0.55 Read average G+C 0.38 Stoichiometry Read depth Lepto III (1X) 3 10 Lepto II (3X) Bacterial Archaeal Fer 1 (1X) G-plasma (1X) Fer 2 (3X) 3 10
0.55 Read average G+C 0.38 Lepto III 3 10 Lepto II Other sampled genomes at low depth (including eukaryotes) 15% of reads Bacterial Archaeal Fer 1 G-plasma Fer 2 3 10
Similarity to Fer1(isolate) to Sequence in Community 78.2% 64.9% Mixed Community Reads 98-100% Fer2 Fer1 G plasma Number of reads .50 .60 .70 .80 .90 1. %id to cultivated Fer1 isolate
Conclusions So Far • The stochiometry of organisms encouraging for the assembly of individual genomes • Assemblies support 16S studies suggesting limited diversity • Isolated Fer1 genome sequences matches genome in environmental sample
How do you know you’ve done it right?Check pair ends against scaffold How do we know that our assembly is correct? At the gross level: check pairs (expect few % due to failing/chimeric clones) Align all reads back against assembled scaffolds scaffolds end where there is no clone coverage in 3kb plasmids Identifies potentially repetitive areas and/or rearrangements
Fer2 vs. fer1 shows local synteny • Fer1 and • Fer2 have avg. nt identity of 78% Fer2 gene on contig Fer1 gene on contig
What does it mean to assemble a community genome? Sample derived from millions of genomes. ? • What is a “species” in the enviroment? • Members of the same species • significantly different (many lineages survive and diverge) • highly similar (selective sweeps)
What does it mean to assemble a community genome? Lepto II : 1 nucleotide variation / 3,000 bp Fer II: 2.2 nucleotide variation / 100 bp
5 Reads of the Same Sequence from 5 Different Members of the Same Species (FerII) 1 1 2 2 4 4 5 5 • CONSENSUS 130953 gtttatattaaatccattgatttctaagcttccggttcttcttccgtataatggagattt 131012 • XYG46314.b1 162 A.......C........................A...........A.............. 103 • XYG44123.b1 673 A.......C........................A...........A.............. 732 • XYG44918.b1 48 A.......C........................A........... 4 • XYG13291.g3 2 .......... 11 • XYG40116.g1 192 ......G..................................................... 133 • XYG3051.b2 396 ......G..................................................... 455 • CONSENSUS 131013 atagcttaataattcatcctccatcatacttatgcttgaacctgataatattatgtatag 131072 • XYG46314.b1 102 ............................................................ 43 • XYG44123.b1 733 ............................................................ 792 • XYG13291.g3 12 ............................................................ 71 • XYG40116.g1 132 ...A........................................................ 73 • XYG3051.b2 456 ...A........................................................ 515 • CONSENSUS 131073 ccttgtagtatccattaattcatcaaatattttctgcattatagatataataccatggtt 131132 • XYG46314.b1 42 .......................................... 1 • XYG44123.b1 793 ........................ 816 • XYG13291.g3 72 ............................................................ 131 • XYG40116.g1 72 T............G....C....................A.................... 13 • XYG3051.b2 516 T............G....C....................A.................... 575 1 1 3 3
Two Haplotypes Among the 5 Different Members of the Same Species (FerII) 1 1 2 2 4 4 5 5 • CONSENSUS 130953 gtttatattaaatccattgatttctaagcttccggttcttcttccgtataatggagattt 131012 • XYG46314.b1 162 A.......C........................A...........A.............. 103 • XYG44123.b1 673 A.......C........................A...........A.............. 732 • XYG44918.b1 48 A.......C........................A........... 4 • XYG13291.g3 2 .......... 11 • XYG40116.g1 192 ......G..................................................... 133 • XYG3051.b2 396 ......G..................................................... 455 • CONSENSUS 131013 atagcttaataattcatcctccatcatacttatgcttgaacctgataatattatgtatag 131072 • XYG46314.b1 102 ............................................................ 43 • XYG44123.b1 733 ............................................................ 792 • XYG13291.g3 12 ............................................................ 71 • XYG40116.g1 132 ...A........................................................ 73 • XYG3051.b2 456 ...A........................................................ 515 • CONSENSUS 131073 ccttgtagtatccattaattcatcaaatattttctgcattatagatataataccatggtt 131132 • XYG46314.b1 42 .......................................... 1 • XYG44123.b1 793 ........................ 816 • XYG13291.g3 72 ............................................................ 131 • XYG40116.g1 72 T............G....C....................A.................... 13 • XYG3051.b2 516 T............G....C....................A.................... 575 1 1 3 3
Two haplotypes Among the 5 Different Members of the Same Species (Fer II) 1 1 2 2 4 4 5 5 • CONSENSUS 130953 gtttatattaaatccattgatttctaagcttccggttcttcttccgtataatggagattt 131012 • XYG46314.b1 162 A.......C........................A...........A.............. 103 • XYG44123.b1 673 A.......C........................A...........A.............. 732 • XYG44918.b1 48 A.......C........................A........... 4 • XYG13291.g3 2 .......... 11 • XYG40116.g1 192 ......G..................................................... 133 • XYG3051.b2 396 ......G..................................................... 455 • CONSENSUS 131013 atagcttaataattcatcctccatcatacttatgcttgaacctgataatattatgtatag 131072 • XYG46314.b1 102 ............................................................ 43 • XYG44123.b1 733 ............................................................ 792 • XYG13291.g3 12 ............................................................ 71 • XYG40116.g1 132 ...A........................................................ 73 • XYG3051.b2 456 ...A........................................................ 515 • CONSENSUS 131073 ccttgtagtatccattaattcatcaaatattttctgcattatagatataataccatggtt 131132 • XYG46314.b1 42 .......................................... 1 • XYG44123.b1 793 ........................ 816 • XYG13291.g3 72 ............................................................ 131 • XYG40116.g1 72 T............G....C....................A.................... 13 • XYG3051.b2 516 T............G....C....................A.................... 575 1 1 3 3
Polymorphisms occur in blocks % polymorphic sites • Long quiet regions separate highly variable segments • Variation is found in blocks of 5-10 genes Local depth ORFs
Summary of Iron Mountain Biofilm • Limited number of predominant species present in biofilm the majority have never been cultured • Several lines of evidence suggest that we can assemble genomes of these organisms • Simplicity of community suggests removal of most variants by natural selection • Now studying the metabolic capabilities of microbes