Comprehensive Analysis of Genomic Sequences Using Multiple Bioinformatics Tools
This project involves the systematic analysis of genomic sequences through the integration of various bioinformatics tools designed for motif discovery, phylogenetic analysis, and statistical evaluations. Key components include AlignAce for alignment, Gibbs for motif detection, and Weeder for sequence analysis. The results are aggregated and analyzed, capturing metrics such as true positives and false positives, to enhance genomic research. The workflow facilitates high-throughput processing and can be customized based on user inputs, including genomic names and specific parameters.


Comprehensive Analysis of Genomic Sequences Using Multiple Bioinformatics Tools
E N D
Presentation Transcript
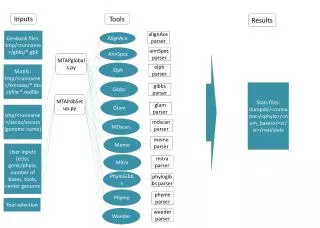
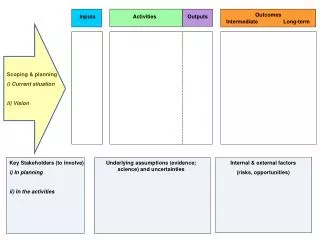
Inputs Tools Results Genbank files: tmp/<runname>/gbks/*.gbk alignAce parser AlignAce AnnSpec annSpec parser MTAPglobals.py Elph elph parser Motifs: tmp/<runname>/kmraws/*.blackfile *.redfile Gibbs Stats files: dumpdir/<runname>/<phylo>/<num_bases>/<cr/sr>/real/stats gibbs parser MTAPdbSetup.py Glam glam parser tmp/<runname>/accoc/accocs (genome name) MDscan mdscan parser Meme meme parser User Inputs (cr/sr, genic/phylo, number of bases, tools, center genome Mitra mitra parser PhyloGibbs phylogibbs parser Phyme phyme parser Tool selection Weeder weeder parser
Inputs Tools Results Sequence to be analyzed alignAce parser AlignAce AnnSpec annSpec parser MTAPglobals.py Elph elph parser Motifs Gibbs Stats files: (number of true positives, true negatives, and false positives) gibbs parser MTAPdbSetup.py Glam glam parser Name of genome to be analyzed MDscan mdscan parser Meme meme parser User Inputs (genic/phylo, number of bases, tools) Mitra mitra parser PhyloGibbs phylogibbs parser Phyme phyme parser Tool selection Weeder weeder parser