Restriction Enzyme Digestion
130 likes | 216 Vues
Explore the world of restriction enzymes and their pivotal role in DNA technology, from cutting specific sequences to creating DNA fingerprints. Learn about the different types and uses of restriction endonucleases, as well as practical techniques like gel electrophoresis. Discover how enzymes like DNA ligase facilitate the creation of recombinant DNA. Expand your knowledge and delve into the fascinating realm of genetic manipulation.

Restriction Enzyme Digestion
E N D
Presentation Transcript
Restriction Enzyme Digestion Timothy G. Standish, Ph. D.
Enzymes are the Tools of DNA Technology • The tools of DNA technology are the same enzymes used by cells to modify their own DNA: • DNA Polymerases • DNA Ligase • One special class of enzyme is pivotal to the cloning of DNA and many other techniques used in DNA Technology • These enzymes are the restriction endonucleases • Restriction - Because for the way they work, they restrict bacteriophages to only one host bacterial strain. They are also restricted to acting on only specific DNA sequences • Endonuclease - They cut nucleic acids in the middle, not just the ends
Restriction Endonucleases • There are a number of different subclasses of restriction endonucleases • Type I - Recognize specific sequences and cut DNA at a nonspecific site > than 1,000 bp away • Type II - Recognize palindromic sequences and cut within the palindrome • Type III - Recognize specific 5-7 bp sequences and cut 24-27 bp downstream of the site. • Type II restriction endonucleases are the most useful class as they recognize specific palindromic sequences in DNA and cut the sugar phosphate backbone within the palindrome
What is a Palindrome? • A palindrome is anything that reads the same forwards and backwards: • English palindromes: • Mom • Dad • Tarzan raised Desi Arnaz rat. • Able was I ere I saw Elba (supposedly said by Napoleon) • Doc note I dissent, a fast never prevents a fatness, I diet on cod.
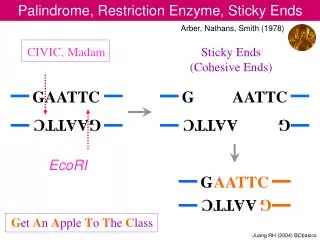
DNA Palindromes • Because DNA is double stranded and the strands run antiparallel, palindromes are defined as any double-stranded DNA in which reading 5’ to 3’ both are the same • Some examples: • The EcoRI cutting site: • 5'-GAATTC-3' • 3'-CTTAAG-5' • The HindIII cutting site: • 5'-AAGCTT-3' • 3'-TTCGAA-5'
Uses of Restriction Endonucleases • Because restriction endonucleases cut specific sequences they can be used to make “DNA fingerprints” of different samples of DNA. As long as the cutting site changes on the DNA or the distance between cutting sites changes, fragments of different sizes will be made. • Because Type II restriction endonucleases cut only at palindromes, they leave “sticky ends” that will base pair with any other fragment of DNA cut with the same enzyme. This is useful in cloning.
EcoRI EcoRI GAATTC CTTAAG GAATTC CTTAAG 1 Digestion AATTC G G CTTAA 2 Annealing of sticky ends Ligase AATTC G G CTTAA 4 Recombinant DNA AATTC G G CTTAA 3 Ligation R. E.s and DNA Ligase Can be used to make recombinant DNA
Gel Electrophoresis • Separates DNA (or RNA or Protein) fragments on the basis of charge and size • Because DNA is an acid, it loses protons in basic buffers, thus it has a negative charge that should be uniform per unit length • Agarose (a polysaccharide) or other gel matrices are difficult for large DNA fragments to move through • The larger the fragment, the more difficulty it has moving through gels • By placing DNA in a gel, then applying a voltage across the gel, the negatively charged DNA will move toward the positive pole • Large fragments lag behind while small fragments move through the gel relatively rapidly
Wells Gel Electrophoresis
Wells Gel Electrophoresis
- Wells Direction of DNA Travel + Gel Electrophoresis Large Small
The End