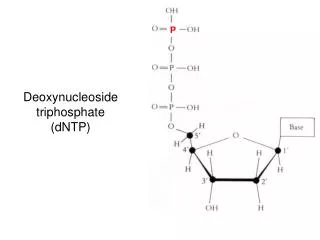
Nucleotide biosynthesis
Nucleotide biosynthesis. Base Nucleoside Nucleotide (base + ribose) (nucleoside + phosphate) A denine A denosine AMP (adenylic acid or adenosine monophosphate) G uanine G uanosine GMP (guanylic acid


Nucleotide biosynthesis
E N D
Presentation Transcript
Nucleotide biosynthesis BaseNucleoside Nucleotide (base + ribose) (nucleoside + phosphate) Adenine Adenosine AMP (adenylic acid or adenosine monophosphate) Guanine Guanosine GMP (guanylic acid or guanosine monophosphate) Uracil Uridine UMP (uridylic acid or uridine monophosphate) Thymine Thymidine TMP (thymidylic acid or thymidine monosphosphate) Cytosine Cytidine CMP (cytidylic acid or cytidine monophosphate)
The purine ring numbering system is: Purines Adenine is 6-aminopurine. Guanine is 2-amino-6-oxopurine.
Purine biosynthesis was investigated by John Buchanan (University of Pennsylvania and MIT) and Robert Greenberg (University of Michigan) in the 1940s. Buchanan fed labelled precursors to pigeons and analysed their incorporation into urate. Birds excrete excess nitrogen as urate (a purine).
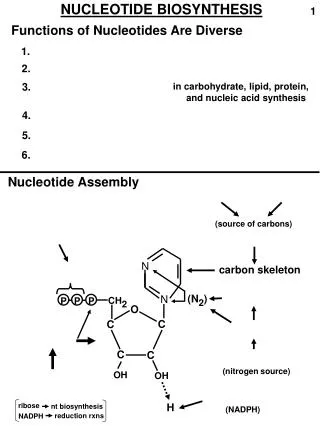
The sequence of precursors that contribute ring atoms is: Glutamine Glycine Formyl-THF Glutamine CO2 Aspartate Formyl-THF All contribute one ring atom except glycine which contributes 3. The purine ring is built up on a ribose sugar that is derived fromPRPP.
6 5 7 N N 1 8 2 N N 4 9 3 The order in which ring atoms are added is: 9 4 5 7 8 3 6 1 2 Q G For Q CO2 D For Ribose-P
Azaserine is a toxic analogue of glutamine that inhibits purine biosynthesis. Glutamine Azaserine Sulfonamide antibiotics inhibit folate biosynthesis and block purine biosynthesis. They cause bacteria to accumulate a biosynthetic intermediate.
Q G For Q CO2 D For 9 4 5 7 8 3 6 1 2 Q 9 G 457 For 8 Q 3 CO2 6 D 1 For 2
Four ATP molecules are used up during construction of each IMP,mostly for activation steps. The intermediate 5-amino imidazole 4-carboxamide ribonucleotide is equivalent to the remains of AMP after the N1-C2 fragment is used for HIS biosynthesis.
IMP is converted to AMP or GMP IMP AMP XMP GMP
Glutamine is the source of amino groups for conversion of XMP to GMP. GTP is required for conversion of IMP to AMP. ATP is required for conversion of IMP to GMP. This helps to balance the concentrations of AMP and GMP in cells.