Network Analysis
Network Analysis. Group Meeting Talk Aidi Stan, Feb 28. Network Analysis. Any type of analysis that involves network information ( pathway ), including Regulatory analysis find controllers ( Network controllability) Pathway analysis summarize and compare Other analysis

Network Analysis
E N D
Presentation Transcript
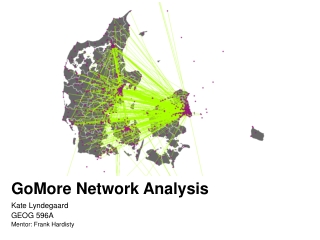
Network Analysis Group Meeting Talk Aidi Stan, Feb 28
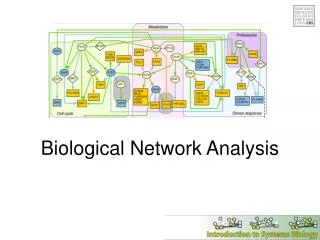
Network Analysis • Any type of analysis that involves network information (pathway), including • Regulatory analysis • find controllers (Network controllability) • Pathway analysis • summarize and compare • Other analysis • predict gene function • find new pathway members • identify functional modules (new pathways)
Pathway Analysis DAVID GSEA Reactome FI Khatri et.al. PLoScomp.bio. 2012
Other analysis • Predict gene function • disease gene: Cipher • drug target: drugCipher • Identify functional modules • ComCipher (co-module) • ClustEx 2.0
Things Worth Noticing • Multilayer network (data integration) • Dynamics • Context (cell type, developmental stage)
Integrated Data Integrated genomic analyses of ovarian carcinoma The Cancer Genome Atlas Research Network. Nature. 2011 • Separate analyzed different data • Used PARADIGM (Bioinformatics. 2010)
Comprehensive molecular portraits of human breast tumours The Cancer Genome Atlas Research Network. Nature. 2012 • Clustering mRNA – expression -> Subtype • mRNA-subtype - miRNA-lusters overlap • Mutations – subtype associations
Integrated Analyses Identify a Master MicroRNA Regulatory Network for the Mesenchymal Subtype in Serous Ovarian Cancer Yang et.al. Cell. 2013 Main steps: • significantly overexpressed genes (2942) • genes correlated with copy number alteration (CNA), DNA methylation, or associated miRNA expression (259) • Among miRNAs which regulate remained genes, three have the largest number or targets (including miR-506) Features: • Only used miRNA-regulatory network • No computational framework
Dynamics Dynamic regulatory network controlling TH17 cell differentiation Yosef et.al. Nature. 2013 Motivation • The molecular circuits that control the differentiation of naive T cells remain largely unknown • Recent studies that reconstructed regulatory networks in mammalian cells have focused on short-term responses Contribution • Combine transcriptional profiling at high temporal resolution, novel computational algorithms, and innovative nanowire-based perturbation tools to systematically derive and experimentally validate a model of the dynamic regulatory network that controls the differentiation of mouse TH17 cells
18 times points along a 72-h time course similar within each phase induction; onset of phenotype and amplification; stabilization and IL-23 signalling
Hypothesis: each of the clusters encompasses genes that share regulators active in the relevant time points • Connected a regulator to a gene from its set of putative targets only if there was also a significant overlap between the regulator’s putative targets and that gene’s cluster • Labelled an edge between a regulator and a gene with a time stamp when the target one is regulated and the regulator node is sufficiently expressed
Other Network analysis of GWAS data (Review) Leiserson et.al. Genetics & Development. 2013 • GWAS-identified SNPs are generally not located in the gene(s) underlying the phenotype of interest (Topology) • GWAS-detected variants do not explain most of the genetic effects found in affected individuals (Network)
Network-based Analysis of Genome Wide Association Data Provides Novel Candidate Genes for Lipid and Lipoprotein Traits Barabasi lab. The American Society for Biochemistry and Molecular Biology. 2013 Steps: • Mapping SNPs to genes. • Construction of a human interactome • Identification of candidate genes using molecular triangulation (MT). • Identification of modules using the jActiveModulemethod (jAM). • Selection of phenotypically coherent (GCM) modules of seed and candidate genes using comorbidity analyses. • Validation and comparison to other methods (CANDID and MetaRanker). • Selection of SNPs representing GCM candidate genes for genotyping in the MDC-CC. Context
Network-based differential gene expression analysis suggests cell cycle related genes regulated by E2F1 underlie the molecular difference between smoker and non-smoker lung adenocarcinoma Wu et.al. BMC Bioinformatics. 2013 Traditional: Differential gene expression Present work: Combine gene-gene co-expression and corresponding expression level (turned into network analysis)