Protein structure Classification
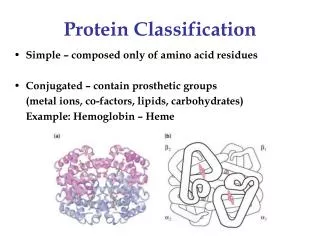
Protein structure Classification. Ole Lund, Associate professor, CBS, DTU. Why classify proteins. Number of solved structures grow rapidly Generate overview of structure types Detect similarities (evolutionary relationships) Set up prediction benchmarks. Classification schemes. SCOP

Protein structure Classification
E N D
Presentation Transcript
Protein structure Classification Ole Lund, Associate professor, CBS, DTU.
Why classify proteins • Number of solved structures grow rapidly • Generate overview of structure types • Detect similarities (evolutionary relationships) • Set up prediction benchmarks OL
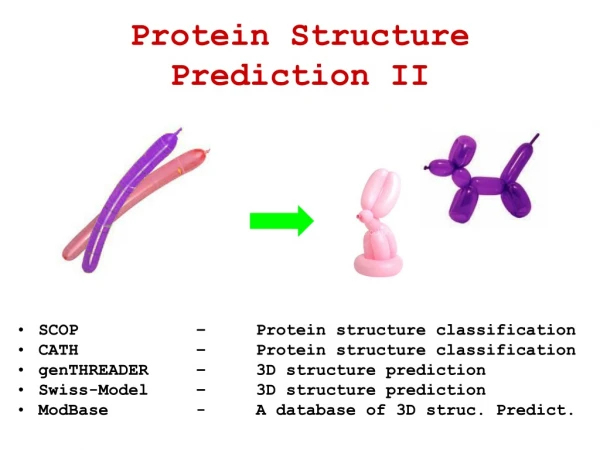
Classification schemes • SCOP • Manual classification (A Murzin) • CATH • Semi manual classification (C orengo) • FSSP • Automatic classification (L Holm) OL
Levels in SCOP • Class 10 • Folds 648 • Superfamilies 1007 • Families 1699 Murzin et al., 1995 http://scop.mrc-lmb.cam.ac.uk/scop/ OL
Major classes in scop • Classes • All alpha proteins • Alpha and beta proteins (a/b) • Alpha and beta proteins (a+b) • Multi-domain proteins • Membrane and cell surface proteins • Small proteins OL
Folds* • Each Class may be divided into one or more folds • Proteins which have the same (>~50%) secondary structure elements arranged the in the same order in the protein chain and in three dimensions are classified as having the same fold *confusingly also called fold classes OL
Superfamilies • Superfamilies are a subdivisions of folds • A superfamily contains proteins which are thought to be evolutionarily related due to • Sequence • Function • Special structural features • Relationships between members of a superfamily may not be readily recognizable from the sequence alone OL
Families • Subdivision of supefamilies • Contains members whose relationship is readily recognizable from the sequence (>~25% sequence identity) • Families are further subdivided in to Proteins • Proteins are divided into Species • The same protein may be found in several species OL
Families OL
CATH • Levels • Class • Architecture • This level is unique to CATH (~30 is defined) • Topology • ~Fold(/superfamily) in SCOP • Homologous Superfamily • ~Superfamily(/family) in SCOP http://www.biochem.ucl.ac.uk/bsm/cath_new/index.html OL
Architecture • Same overall arrangement of secondary structures • Example: The architecture :Two layer beta sheet proteins contains different folds each with a distinct number and connectivity of strands OL
FSSP • Fully automated classification • Automatic update • Database contains structural alignments • Tree of protein structures OL
Links • PDB • www.rcsb.org/pdb/ • SCOP • scop.mrc-lmb.cam.ac.uk/scop/data/scop.b.html • CATH • www.biochem.ucl.ac.uk/bsm/cath_new/index.html • FSSP • http://www.bioinfo.biocenter.helsinki.fi:8080/dali/index.html OL