Genetic Diversity Analysis Using UPGMA Tree Based on Allelic Profiles of Various STs
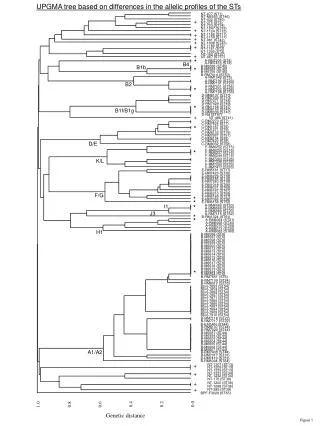
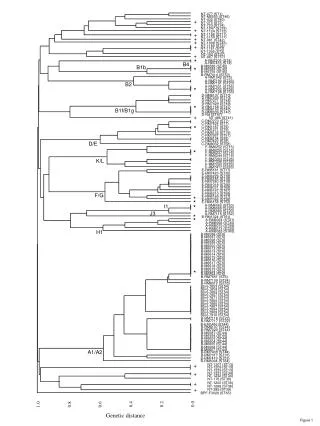
This research employs the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) to construct a phylogenetic tree. The study analyzes genetic relationships by assessing allele differences among various sequence types (STs), including NT-477 (ST1), NT-M6593 (ST46), and others. This representation aids in visualizing genetic diversity and evolutionary relationships within populations, providing insights into the structure of genetic variation across different STs.

Genetic Diversity Analysis Using UPGMA Tree Based on Allelic Profiles of Various STs
E N D
Presentation Transcript
UPGMA tree based on differences in the allelic profiles of the STs NT-477 (ST1) NT-M6593 (ST46) + NT-432 (ST40) NT-375 (ST3) NT-723 (ST14) + NT-1247 (ST33) NT-1124 (ST12) + NT-1158 (ST11) + NT-1159 (ST11) + NT-981 (ST42) NT-1008 (ST43) + NT-1180 (ST2) NT-1181 (ST2) NT-1292 (ST2) + NT-162 (ST37) NT-667 (ST57) * A-RM7205 (ST4) B4 A-RM7416 (ST4) B1b * B-M6588 (ST45) B-M6589 (ST45) B-M6755 (ST45) B-RM7414 (ST50) A-RM1042 (ST5) A-RM7190 (ST23) B2 A-RM7191 (ST23) * A-RM7031 (ST56) A-RM7032 (ST56) A-RM7198 (ST56) D-RM6137 (ST10) D-RM7033 (ST10) D-RM7271 (ST48) D-RM7429 (ST49) * D-RM1168 (ST47) B1f/B1g D-RM6150 (ST47) D-RM8039 (ST47) + D-Rd (ST47) NT-486 (ST41) C-RM7270 (ST7) C-RM7424 (ST7) * C-RM1167 (ST9) C-RM1271 (ST9) C-RM6132 (ST19) C-RM7267 (ST51) C-RM6134 (ST8) D/E C-RM7422 (ST8) C-RM8032 (ST58) F-RM6252 (ST15) * F-RM6255 (ST16) F-RM6237 (ST16) F-RM6244 (ST16) K/L F-RM7283 (ST26) F-RM7298 (ST26) F-RM7290 (ST29) F-RM7417 (ST63) E-RM6181 (ST17) E-RM7423 (ST32) E-RM6229 (ST18) E-RM6185 (ST18) E-RM7280 (ST18) E-RM1018 (ST66) E-RM8031 (ST69) E-RM6167 (ST67) F/G E-RM6210 (ST68) * E-RM6169 (ST27) * E-RM7066 (ST28) E-RM6158 (ST52) I1 * A-RM6062 (ST20) A-RM6069 (ST20) J3 A-RM6080 (ST20) * A-RM7115 (ST62) * B-RM1324 (ST61) A-RM6064 (ST21) A-RM6068 (ST30) A-RM6070 (ST59) A-RM6073 (ST25) H1 A-RM6083 (ST60) B-M6596 (ST6) B-M6597 (ST6) B-M6598 (ST6) B-M6599 (ST6) B-M6600 (ST6) B-M6612 (ST6) B-M6613 (ST6) B-M6614 (ST6) B-M6615 (ST6) B-M6616 (ST6) B-M6617 (ST6) B-M6618 (ST6) * B-M6619 (ST6) B-M6624 (ST6) B-M6894 (ST6) B-RM7651 (ST6) B-RM7109 (ST24) B-RM8012 (ST55) B(iv)-7853 (ST54) B(iv)-7854 (ST54) B(iv)-7863 (ST54) B(iv)-7867 (ST54) B(iv)-7871 (ST54) B(iv)-7884 (ST54) B(iv)-7885 (ST54) B(iv)-7887 (ST54) B(iv)-7894 (ST54) B(iv)-7909 (ST54) B(iv)-7910 (ST54) B-RM7118 (ST22) B-RM7717 (ST22) B-EAGAN (ST44) B-RM6107 (ST44) B-RM7020 (ST44) B-M6587 (ST44) B-M6594 (ST44) B-M6603 (ST44) B-M6604 (ST44) B-M6605 (ST44) B-M6606 (ST44) A1/A2 B-M6607 (ST44) B-RM7430 (ST44) B-RM7017 (ST31) B-RM7419 (ST53) B-RM6094 (ST64) + NT-1207 (ST13) NT-1209 (ST13) NT-1233 (ST13) + NT-1231 (ST34) NT-1232 (ST34) NT-176 (ST38) + NT-1200 (ST36) NT-1268 (ST36) + NT-285 (ST39) BPF-F3028 (ST65) 0.0 0.2 0.4 1.0 0.6 0.8 Genetic distance Figure 1