Recombinant DNA Technology - The Basis & Applications
350 likes | 417 Vues
Explore the principles and applications of DNA recombinant technology in genetic engineering. Learn about gene manipulation, cloning, gene expression, and purification processes for various treatments. Enhance your knowledge with diverse tools and techniques.

Recombinant DNA Technology - The Basis & Applications
E N D
Presentation Transcript
DNA recombinant Genetic Engineering The manipulation of an organism endowment by introducing or eliminating specific gene A gene of interest is inserted into another organism, enabling it to be cloned, and thus studied more effectively Design and construction of new combinations of genes (DNA) New combinations/arrangements of DNA DNA cloning
DNA Recombinant Technology Technology used in the isolation or synthesis and joining together of unlike pieces of DNA These recombinant DNA molecules can then be introduced into bacteria, yeasts, or other cells where they can replicate and function (code for protein synthesis)
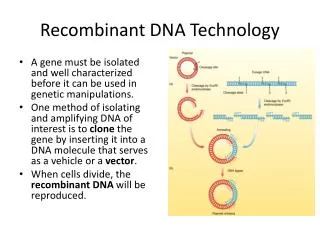
Overview of Genetic Engineering • Gene of interest is isolated from appropriate organism • Gene is recombined with a vector (carrier) DNA molecule • Recombinant DNA is introduced into appropriate host cell • Recombinant DNA is expressed at high levels in host cell • Gene product may be purified for use in treatments (antibiotics, hormones, etc.)
Why Detailed studies of the structure and function of a gene at the molecular level require large quantities of the individual gene in pure form
Tools Restriction and ligation enzymes Plasmid as a vector Host Cells
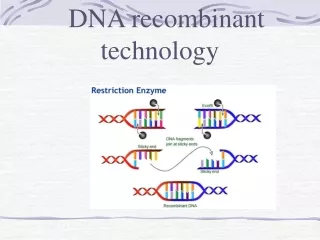
Restriction Enzymes Molecular scissors which isolated from bacteria where they are used as Bacterial defense against viruses Molecular scalpels to cut DNA in a precise and predictable manner Enzyme produced by bacteria that typically recognize specific 4-8 base pair sequences called restriction sites, and then cleave both DNA strands at this site A class of endo-nucleases that cleavage DNA after recognizing a specific sequence Members of the class of nucleases
Nuclease Breaking the phosphodiester bonds that link adjacent nucleotides in DNA and RNA molecules • Endonuclease • Cleave nucleic acids at internal position • Exonuclease • Progressively digest from the ends of the nucleic acid molecules
Restriction Enzymes • There are already more than 1200 type II enzymes isolated from prokaryotic organism • They recognize more than 130 different nucleotide sequence • They scan a DNA molecule, stopping only when it recognizes a specific sequence of nucleotides that are composed of symetrical, palindromic sequence • Palindromic sequence: • The sequence read forward on one DNA strand is identical to the sequence read in the opposite direction on the complementary strand • To Avoid confusion, restriction endo-nucleases are named according to the following nomenclature
Nomenclature • The first letter is the initial letter of the genus name of the organism from which the enzyme is isolated • The second and third letters are usually the initial letters of the organisms species name. It is written in italic • A fourth letter, if any, indicates a particular strain organism • Originally, roman numerals were meant to indicate the order in which enzymes, isolated from the same organisms and strain, are eluted from a chromatography column. More often, the roman numerals indicate the order of discovery
Restriction enzymes Restriction enzymes can be grouped by: • number of nucleotides recognized (4, 6,8 base-cutters most common) • kind of ends produced (5’ or 3’ overhang (cohesive=sticky), blunt=flush) • degenerate or specific sequences • whether cleavage occurs within the recognition sequence Become familiar with the back of your molecular biology catalog!
A restriction enzyme (EcoRI) 1. 6-base cutter 2. Specific palindromic sequence (5’GAATTC) 3. Cuts within the recognition sequence (type II enzyme) 4. produces a 5’ overhang (sticky end)
Cloning A collection of molecules or cells, all identical to an original molecule or cell • To "clone a gene" is to make many copies of it - for example, in a population of bacteria • Gene can be an exact copy of a natural gene • Gene can be an altered version of a natural gene • Recombinant DNA technology makes it possible
Vector an autonomously replicating genetic element used to carry DNA fragments into a host, typically E. coli, for the purpose of gene cloning • Plasmid vector • Bacteriophage gamma vector
Plasmids Naturally occurring extra-chromosomal DNA • Plasmids are circular double stranded DNA • Plasmids can be cleaved by restriction enzymes, leaving sticky ends • Artificial plasmids can be constructed by linking new DNA fragments to the sticky ends of plasmid
Cloning Vectors Plasmids that can be modified to carry new genes • Plasmids useful as cloning vectors must have • a replicator (origin of replication) • a selectable marker (antibiotic resistance gene) • a cloning site (site where insertion of foreign DNA will not disrupt replication or inactivate essential markers
Chimeric Plasmids Named for mythological beasts with body parts from several creatures • After cleavage of a plasmid with a restriction enzyme, a foreign DNA fragment can be inserted • Ends of the plasmid/fragment are closed to form a "recombinant plasmid" • Plasmid can replicate when placed in a suitable bacterial host
Directional Cloning Often one desires to insert foreign DNA in a particular orientation • This can be done by making two cleavages with two different restriction enzymes • Construct foreign DNA with same two restriction enzymes • Foreign DNA can only be inserted in one direction
Host Cells • Propagation of a DNA sequence must take place inside a living cell (host cells) • Eschericia coli: • It provides a relatively simple and well understood genetic environment • The way to isolate plasmid is understood • It contains a single chromosome of approximately 5 Mbp • The genetic code is nearly universal • It replicates once every 22 minutes • It grows best with incubation at 37°Cin a culture medium that approximately the nutrient available in the human digestive tract
Bacterial transformation • The cellular uptake and expression of DNA in a bacteria • Introduction of DNA into competent cell of bacteria • Requested element in transformation: • A suitable host organism in which to insert the gene • A self-replicating vector to carry the gene into the host organism • A means of selection for host cells that have taken up the gene
Selection of Transformant A particularly important selective advantage offered by plasmid is antibiotic resistance gene It encodes for proteins that disable antibiotics secreted by microorganism with which bacteria compete Antibiotics function by several different mechanism Antibiotics resistance: A selectable marker that allows one to positively identify cells that have been induced to take up plasmid DNA Penicillin family (including ampicillin) interfere with cell wall biosynthesis Kanamycin, tetracyclin, and chloramphenicol arrest bacterial cell growth by blocking various steps in protein synthesis
Protein expression • - Gene is inserted into plasmid • - Plasmid is transformed into a host • cell (E. coli) • Cell culture is prepared • Each cell contains several copies of the plasmid with gene • Gene expression leads to the production of protein • Protein level may reach 30% of • total cellular protein • Isolation of protein